Research
, Volume: 23( 1) DOI: 10.37532/0974-7419.2022.23(1).233.Rapid determination of PCDDs, PCDFs and DL-PCBs in foods, feedingstuffs and vegetable oils using new modified acid silica
- *Correspondence:
- Ovanes Chakoyan Fidelitas Lab, Department of Chromatography, Varna, Bulgaria., E-mail: ovanes.chakoyan@fidelitasgroup.com
Received: December 08, 2023, Manuscript No. tsac-22-82219; Editor assigned: December 11, 2023, PreQC No. tsac-22-82219 (PQ); Reviewed: December 11, 2023, QC Notsac-22-82219 (Q); Revised: December 13, 2023, Manuscript No tsac-22-82219 (R); Published date: Januray 27, 2023. DOI:10.37532/0974-7419.2023.23(1).233.
Citation:Chakoyan O. Rapid determination of PCDDs, PCDFs and DL-PCBs in foods, feedingstuffs and vegetable oils using new modified acid silica. Anal Chem Ind J. 2023;23(1):233.
Abstract
Polychlorinated dibenzo-p-dioxins (PCDDs), dibenzofurans (PCDFs) and dioxin-like polychlorinated biphenyls (DL-PCBs) are persistent organic pollutants (POPs), which in the recent years received huge attention due to their extreme stability, high potential toxicity and bioaccumulation in food chains. Main source of human exposure to these compounds is discovered in foods of animal origin, especially foods rich of animal fat. The target of the present study was to set up an analytical method for the determination of PCDDs/PCDFs and DL-PCB in vegetable oils, sunflower meals, sunflower seeds, rapeseeds and milk powder. The first step consisted of a semi-automatic Soxhlet extraction for 3 hours, by using a mixture of Hexane: Acetone - 80: 20, followed by acid digestion with 55% acid silica, and filtration. After concentration, the extract is purified on multilayer column (silica gel, silica-KOH, silica-H2SO4 anhydrous Na2SO4) followed by an alumina column separation in two fractions (first fraction containing PCDDs/PCDFs, and the second containing only DL-PCBs). The purified extract was then analyzed by GC/MS/MS. The newly developed approach in our lab was capable to reduce the overall time of sample preparation down to seven hours/ per sample from initial of what time. Since the method shows good mean recoveries for all labelled congeners spiked in the samples (for PCDDs/PCDFs – 80-110%, for DL-PCBs – 70-85%), we assumed the absence of over estimation or underestimation in the analyzed samples.
Keywords
Oil; Extraction; Polychlorinated-p-dioxins; Polychlorinated dibenzofurans; Polychlorinatedbiphenyls; GC/MS/MS
Introduction
Dioxins are a group of persistent chemicals which are not produced chemically, but are formed during combustion (burning) processes and as by-products of industrial processes. Polychlorinated biphenyls (PCBs), have similar chemical structure as dioxins. They have been used in transformers, building materials, lubricants, coatings, plasticizers and inks, although their use has now largely been phased out. Both the dioxins and the PCBs are highly resistant to breakdown processes, and consequently persist in the environment, followed by uptake into the food chain. Up to 90% of human exposure to dioxin results from the consumption of food containing dioxins, mainly feedstuffs of animal origin with high fat content, since these contaminants accumulated in fatty tissues. Food stuffs in which dioxins can occur includes meat, fish, eggs, and milk.
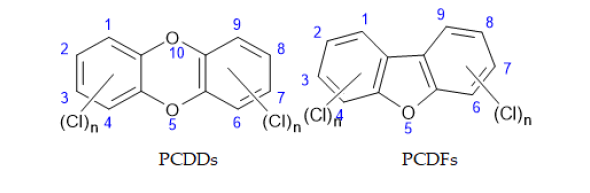
General structure of these compounds represents planar, tricyclic aromatic ether, which can have up to eight chlorine atoms attached to carbon atoms in the benzene rings. The number and position of chlorine atoms on the rings determine different isomers called congeners. Congeners with equal number of chlorine atoms are called homologues, and homologues with different chlorine substitution are called isomers. In general, term dioxins is used to refer to 75 congeners of PCDDs and 135 congeners of PCDFs (total of 210 compounds). Out of these 210 compounds, only 17 congeners (7 PCDDs and 10 PCDFs) with chlorine atoms at position 2, 3, 7 and 8 showing toxicity. FIG.1. shows the general structure of PCDDs and PCDFs.
Polychlorinated biphenyls (PCBs) are nonpolar, chlorinated aromatic hydrocarbons with chlorine atoms from 1 to 10, generating 209 compounds (congeners) as dioxins.
PCBs with equal number of chlorine atoms also called homologues, and homologues with different substitution positions are referred to as isomers. Positions 2, 2’, 6 and 6’ are called ortho-positions; 3, 3’, 5 and 5’ are called meta-positions; and positions 4 and 4’ are para-positions. Rings in biphenyls can be planar or nonplanar, depending of the steric and electronic effects of chlorine atoms, especially in ortho-position, where bulky chlorine atoms forces aromatic benzene rings to rotate out of planar configuration. Meta- and para-PCBs have planar molecules and these compounds are called Coplanar Congeners (co-PCBs)(FIG.2).
When more than two chlorine atoms in ortho-position are present, PCBs assume nonplanar con figuration and are indicated as nonplanar congeners.
Due their lipophilic nature, PCDDs, PCDFs and PCBs are usually found adhering or dissolved in rich of lipid content foof stuffs like meat, fatty fish, milk, dairy products and vegetable oils. Therefore, methods of analysis for official control of the level of lipophilic toxic compounds in EU-regulated foods of animal and plant origin includes an initial lipid extraction step (liquid-liquid extraction, PLE extraction, Soxhlet extraction),which isolates the lipids from potentially interfering compounds such as carbohydrates.
The first step in food preparation methods after extraction is removal of lipids content completely as possible. EPA 8290, EPA 1613 and ISO 16215 recommend uses of concentrated sulphuric acid lipid digestion, and also different ratio impregnated silica gel with sulphuric acid column chromatography. Direct digestion with concentrated mineral acid leads to carbonization of compounds of interest, and therefor to low recoveries of PCDD/PCDF and PCBs.
In this study, we performed a new approach for eliminating lipid interfering compounds, with good recoveries and without complex pre-cleanup procedure with sulphuric acid leading to huge losses and requiring very carefully precautions measures. Using a novel triple quadrupole GC/MS/MS system equipped with highly efficient detector and three forms of noise-reduction technologies. The MS analyzer was equipped with BEIS (Boosted Efficiency Ion Source) that maximizes efficiency through
Materials and Methods
Chemicals and standards
17 native and 13C-labeled PCDDs/PCDFs congeners, 12 native and 13C-labeled DL-PCBs were selected for current study, purchased from Wellington Laboratories (Greyhound Chromatography, UK). For PCDDs/PCDFs and DL-PCBs, a nine-point calibration curve ranging from 0.01 pg/µl to 50 pg/µl and from 0.1 pg/µl to 50 pg/µl were used, respectively. 13C-labeled congeners using isotope dilution method were presents in every calibration solution at concentration 2 pg/µl and 5 pg/µl, accordingly. All standard solutions and final extracts before injection were made of nonane (Fischer Scientific, USA)
All reagents used for the analysis of PCDDs/PCDFs and PCBs were of trace analysis grade. However n-hexane, dichloromethane and acetone were supplied from Sigma-Aldrich and Fischer Scientific respectively. Additional equipment includes nitrogen evaporator (manufacturer and model no), rotatory vacuum evaporator (Buchi R100, Switzerland), an analytical precision scale (RADWAG AS 220.R2, Poland), Laboratory Mill Perten 120, Sweden), Hot extraction unit (FOSS ST243 SOXTEC, Denmark).
Sample collections
Selected sunflower meal, sunflower seed and rapeseed samples were collected at local feed processing plants as for the edible vegetable oils and milk powder samples were collected from grocery stores throughout city Varna. The cereals and cereal bran samples containing barley, wheat and corn were collected from various localities of Bulgaria.
Sample Preparation
Soxhlet extraction of Sunflower meals, cereals, cereals-based foods and milk powder: All samples except for vegetable oils were extracted with (full form and detail of FOSS) FOSS extraction device. Briefly, 10 g of sample was weighted to 0.01 g accuracy and place in extraction cell, spiked with extraction labelled 13C-PCDDs/PCDFs and DL-PCBs congeners, and leave for equilibration around 30 min and extracted with solvent mixture of Hexane and Acetone is 80:20, for 3 hours. After that the extract was evaporated to dryness. After that, sunflower meals and milk powder extract was weighted for determination lipid content. The whole extracts were dissolved in hexane, and added 55% acid silica. The samples were shaken vigorously for 30 sec. and put into centrifugation at 5000 rpm for 5 min, filtrated and evaporated do dryness
Extraction of Sunflower seeds and rapeseeds: Briefly, 10 g milled sunflower seed and rapeseed samples were extracted for 3 hours. After concentration, determination of lipid content was obtained gravimetrically. The extracted fat was fortified with extraction labelled mixtures of 13C - PCDDs/PCDFs and DL-PCBs congeners, and leave for homogenization on around 5 min. The oilseed extracts were directly dissolved in hexane and mixed with 55% acid/silica
Each vegetable oil was weighted around 2.5 g, dissolved in 30 ml hexane and performed acid digestion with 17 g of 55% acid-silica, followed by filtration of clean extract and evaporation to dryness.
Extraction of cereals and cereal-based products: 10 gm milled cereal samples were extracted with solvent mixture Hexane: DCM (1:1) for 3 hours evaporated on rotatory evaporator to dryness and subjected directly to multilayer column cleanup.
Multilayer Column Clean-up: All samples except cereal extract , which are directly put on multilayer column purification after filtration and evaporation to dryness, were subjected to cleanup additionally on multilayer acid/base silica column in following order ( 1 g silica, 5 g NaOH-silica, 10 g 44% acid-silica, 1 g sodium sulfate), with hexane. Separation of dioxins and PCBs was accomplished on alumina column (5 g) by wet filling with hexane. The first fraction containing non-ortho and mono-ortho-PCBs was eluted by 70 ml hexane: DCM (95:5), and the second fraction with PCDDs/PCDFs was eluted with 45 ml Hexane: DCM (50:50). After evaporation to complete dryness, each fraction was quantitavely transferred to vial with insert (300 μl volume), spiked with 13C12-1,2,34-TCDD/ 13C12-1,2,3,7,8,9-HxCDD recovery standard mixture for PCDD/PCDF and 13C-PCB70,PCB 111, PCB138 and PCB 189 for DL- PCBs, concentrated to a 20 μl final volume and injected in GC/MS/MS [1-2](FIG.3).
Instrumentation and Measurements
A Shimadzu GC 8000 series gas chromatograph equipped with triple quadrupole mass detector TQ8050 was used. The MS analyzer was equipped with a BEIS ionization chamber. The injection volume was set to 2 µl for PCDDs/PCDFs and DL-PCBs. Chromatographic separations of calibration standards and all extracts for PCDDs/PCDFs were performed on a SH Rxi-5Sil MS 60m × 0.25 mm I.D. × 0.25 µm( Shimadzu, USA) using a injector temperature at 280°C in split less mode and oven temperature program starting at 150°C (1 min), ramp at 200°C/min until 220°C, 200 C/min until 260°C (3 min), 50°C/min until 320°C (3.5 min) , and total run of 47 min. . Ion source temperature was set to 230°C and interface temperature was set to 300°C. Helium (1.5 ml/min) was used as the carrier gas. The MS analyzer was operated in MRM mode, with collision energy and transitions listed in TABLE.1. For determination of PCDDs/PCDFs [3].
TABLE 1 PCDD/PCDF Masses And Ce
| Compound name | Quantitative ion | Ce | Reference ion | Ce |
|---|---|---|---|---|
| 2,3,7,8-TCDD | 319.9>256.9 | 20 | 321.9>258.9 | 20 |
| 1,2,3,7,8-PCDD | 335.9>292.9 | 20 | 353.9>290.9 | 20 |
| 1,2,3,4,7,8-HXDD | 389.8>326.9 | 22 | 391.8>328.9 | 22 |
| 1,2,3,6,7,8-HXDD | 389.8>326.9 | 22 | 391.8>328.9 | 22 |
| 1,2,3,7,8,9-HXDD | 389.8>326.9 | 22 | 391.8>328.9 | 22 |
| 1,2,3,4,6,7,8-HPDD | 423.8>360.8 | 22 | 425.8>362.8 | 22 |
| OCDD | 457.7>394.7 | 22 | 459.7>396.7 | |
| 2,3,7,8-TCDF | 303.9>240.9 | 28 | 305.9>242.9 | 28 |
| 1,2,3,7,8-PCDF | 355.9>292.9 | 20 | 353.9>290.9 | 20 |
| 2,3,4,7,8-PCDF | 339.9>276.9 | 30 | 337.9>274.9 | 30 |
| 1,2,3,4,7,8-HXDF | 389.8>326.9 | 22 | 391.8>328.9 | 22 |
| 1,2,3,6,7,8-HXDF | 373.8>310.9 | 30 | 375.8>312.9 | 30 |
| 2,3,4,6,7,8-HXDF | 373.8>310.9 | 30 | 375.8>312.9 | 30 |
| 1,2,3,7,8,9-HXDF | 373.8>310.9 | 30 | 375.8>312.9 | 30 |
| 1,2,3,4,6,8,9-HPDF | 419.8>355.9 | 30 | 421.8>357.9 | 30 |
| 1,2,3,4,7,8,9-HPDF | 407.8>344.8 | 30 | 409.8>346.8 | 30 |
| OCDF | 441.8>378.8 | 30 | 443.8>380.8 | 30 |
| 2,3,7,8-TCDD( 13C12) | 315.9>251.9 | 28 | 317.9>253.9 | 28 |
| 1,2,3,7,8-PCDD( 13C12) | 367.9>303.9 | 20 | 365.9>301.9 | 20 |
| 1,2,3,4,7,8-HXDD( 13C12) | 401.8>337.9 | 22 | 399.9>335.9 | 22 |
| 1,2,3,6,7,8-HXDD( 13C12) | 401.8>337.9 | 22 | 399.9>335.9 | 22 |
| 1,2,3,7,8,9-HXDD( 13C12) | 401.8>337.9 | 22 | 399.9>335.9 | 22 |
| 1,2,3,4,6,7,8-HPDD( 13C12) | 419.8>355.9 | 30 | 421.8>357.9 | 30 |
| OCDD( 13C12) | 469.8>405.8 | 22 | 471.8>407.8 | 22 |
| 2,3,7,8-TCDF( 13C12) ( 13C12) | 315.9>251.9 | 28 | 317.9>253.9 | 28 |
| 1,2,3,7,8-PCDF( 13C12) ( 13C12) | 351.9>287.9 | 30 | 349.9>285.9 | 30 |
| 2,3,4,7,8-PCDF( 13C12) | 351.9>287.9 | 30 | 349.9>285.9 | 30 |
| 1,2,3,4,7,8-HxDF( 13C12) | 385.8>321.9 | 30 | 387.8>323.9 | 30 |
| 1,2,3,6,7,8-HxDF( 13C12) | 385.8>321.9 | 30 | 387.8>323.9 | 30 |
| 2,3,4,6,7,8-HxDF( 13C12) | 373.8>310.9 | 30 | 375.8>312.9 | 30 |
| 1,2,3,7,8,9-HxDF( 13C12) | 385.8>321.9 | 30 | 387.8>323.9 | 30 |
| 1,2,3,4,6,8,9-HpDF( 13C12) | 419.8>355.9 | 30 | 421.8>357.9 | 30 |
| 1,2,3,4,7,8,9-HpDF( 13C12) | 419.8>355.9 | 30 | 421.8>357.9 | 30 |
| OCDF( 13C12) | 453.8>389.8 | 30 | 455.8>391.8 | 30 |
Separations of DL-PCBs were performed on the same column, injector was set to 2500C and using different oven temperature program starting at 120ºC (1 min), ramp at 200C/min until 180ºC, 50C/min until 200ºC, 2ºC/min until 240ºC (10 min), 2ºC /min until 290ºC (5 min) and total run of 68 min. Ion source temperature was set to 230ºC and interface temperature was set to 300ºC. Helium (2.15 ml/min) was used as the carrier gas. The MS analyzer was operated in MRM mode, with collision energy and transitions listed in TABLE. 2. for determination of DL-PCBs.
TABLE 2. PCBS Masses and Ce
| Compounds name | Quantitative ion | Ce | Reference ion | Ce |
|---|---|---|---|---|
| PCB 77 | 289.9>219.9 | 26 | 291.9>221.9 | 26 |
| PCB 81 | 289.9>219.9 | 26 | 291.9>221.9 | 26 |
| PCB 105 | 323.9>253.9 | 26 | 325.9>255.9 | 26 |
| PCB 114 | 323.9>253.9 | 26 | 325.9>255.9 | 26 |
| PCB 118 | 323.9>253.9 | 26 | 325.9>255.9 | 26 |
| PCB 123 | 323.9>253.9 | 26 | 325.9>255.9 | 26 |
| PCB 126 | 323.9>253.9 | 26 | 325.9>255.9 | 26 |
| PCB 156 | 359.9>289.9 | 28 | 361.9>291.9 | 28 |
| PCB 157 | 359.9>289.9 | 28 | 361.9>291.9 | 28 |
| PCB 167 | 359.9>289.9 | 28 | 361.9>291.9 | 28 |
| PCB 169 | 359.9>289.9 | 28 | 361.9>291.9 | 28 |
| PCB 189 | 393.8>323.9 | 28 | 395.8>325.9 | 28 |
| 13C12-PCB 77 | 301.9>231.9 | 26 | 303.9>233.9 | 26 |
| 13C12-PCB 81 | 301.9>231.9 | 26 | 303.9>233.9 | 26 |
| 13C12-PCB 105 | 335.9>265.9 | 26 | 337.9>267.9 | 26 |
| 13C12-PCB 114 | 335.9>265.9 | 26 | 337.9>267.9 | 26 |
| 13C12-PCB 118 | 335.9>265.9 | 26 | 337.9>267.9 | 26 |
| 13C12-PCB 123 | 335.9>265.9 | 26 | 337.9>267.9 | 26 |
| 13C12-PCB 126 | 335.9>265.9 | 26 | 337.9>267.9 | 26 |
| 13C12-PCB 156 | 371.8>301.9 | 28 | 373.8>303.9 | 28 |
| 13C12-PCB 157 | 371.8>301.9 | 28 | 373.8>303.9 | 28 |
| 13C12-PCB 167 | 371.8>301.9 | 28 | 373.8>303.9 | 28 |
| 13C12-PCB 169 | 371.8>301.9 | 28 | 373.8>303.9 | 28 |
| 13C12-PCB 189 | 405.8>323.9 | 28 | 395.8>325.9 | 28 |
The GC/MS/MS system was calibrated for PCDD.PCDF using response factors generated from a nine-point curve at the level concentrations presented in the TABLE 3 and TABLE 4.
TABLE 3. Calibration solutions of PCDD/PCDF (ng/ml)
| Analyte | Calibration levels | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Native standard | CS1 | CS2 | CS3 | CS4 | CS5 | CS6 | CS7 | CS8 | CS9 |
| 2,3,7,8-TCDD | 0.025 | 0.05 | 0.1 | 0.5 | 2 | 5 | 10 | 20 | 50 |
| 2,3,7,8-ТCDF | 0.025 | 0.05 | 0.1 | 0.5 | 2 | 5 | 10 | 20 | 50 |
| 1,2,3,7,8-PCDD | 0.025 | 0.05 | 0.1 | 0.5 | 2 | 5 | 10 | 20 | 50 |
| 1,2,3,7,8-PCDF | 0.025 | 0.05 | 0.1 | 0.5 | 2 | 5 | 10 | 20 | 50 |
| 2,3,4,7,8-PCDF | 0.025 | 0.05 | 0.1 | 0.5 | 2 | 5 | 10 | 20 | 50 |
| 1,2,3,4,7,8-HXDF | 0.05 | 0.1 | 0.2 | 1 | 4 | 10 | 20 | 40 | 100 |
| 1,2,3,6,7,8-HXDF | 0.05 | 0.1 | 0.2 | 1 | 4 | 10 | 20 | 40 | 100 |
| 2,3,4,6,7,8-HXDF | 0.05 | 0.1 | 0.2 | 1 | 4 | 10 | 20 | 40 | 100 |
| 1,2,3,4,7,8-HXDD | 0.05 | 0.1 | 0.2 | 1 | 4 | 10 | 20 | 40 | 100 |
| 1,2,3,6,7,8-HXDD | 0.05 | 0.1 | 0.2 | 1 | 4 | 10 | 20 | 40 | 100 |
| 1,2,3,7,8,9-HXDD | 0.05 | 0.1 | 0.2 | 1 | 4 | 10 | 20 | 40 | 100 |
| 1,2,3,7,8,9-HXDF | 0.05 | 0.1 | 0.2 | 1 | 4 | 10 | 20 | 40 | 100 |
| 1,2,3,4,6,7,8-HPDF | 0.05 | 0.1 | 0.2 | 1 | 4 | 10 | 20 | 40 | 100 |
| 1,2,3,4,6,7,8-HPDD | 0.05 | 0.1 | 0.2 | 1 | 4 | 10 | 20 | 40 | 100 |
| 1,2,3,4,7,8,9-HPDF | 0.05 | 0.1 | 0.2 | 1 | 4 | 10 | 20 | 40 | 100 |
| OCDD | 0.1 | 0.2 | 0.4 | 2 | 8 | 20 | 40 | 80 | 200 |
| OCDF | 0.1 | 0.2 | 0.4 | 2 | 8 | 20 | 40 | 80 | 200 |
| Internal standards | |||||||||
| 13С12-2,3,7,8-TCDD | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| 13С12-2,3,7,8-ТCDF | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| 13С12-1,2,3,7,8-PCDD | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| 13С12-1,2,3,7,8-PCDF | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| 13С12-2,3,4,7,8-PCDF | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| 13С12-1,2,3,4,7,8-HXDF | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| 13С12-1,2,3,6,7,8-HXDF | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| 13С12-2,3,4,6,7,8-HXDF | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| 13С12-1,2,3,4,7,8-HXDD | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| 13С12-1,2,3,6,7,8-HXDD | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| 13С12-1,2,3,7,8,9-HXDD | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| 13С12-1,2,3,7,8,9-HXDF | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| 13С12-1,2,3,4,6,7,8-HPDF | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| 13С12-1,2,3,4,6,7,8-HPDD | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| 13С12-1,2,3,4,7,8,9-HPDF | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| 13С12-OCDD | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 4 |
| 13С12-OCDF | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 4 | 4 |
| Recovery standards | |||||||||
| 13С12 - 1,2,3,4-TCDD | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| 13С12 -12,3,4,6,9-HXDF | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| 13С12 -1,2,3,4,6,8,9-HPDF | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
TABLE 4. Calibration solutions of DL-PCBs (ng/ml)
| Analyte | Calibration levels | |||||||
|---|---|---|---|---|---|---|---|---|
| Native standard | CS1 | CS2 | CS3 | CS4 | CS5 | CS6 | CS7 | CS8 |
| PCB 77 | 0.2 | 0.5 | 1 | 2 | 5 | 10 | 20 | 50 |
| PCB 81 | 0.2 | 0.5 | 1 | 2 | 5 | 10 | 20 | 50 |
| PCB 105 | 0.2 | 0.5 | 1 | 2 | 5 | 10 | 20 | 50 |
| PCB 114 | 0.2 | 0.5 | 1 | 2 | 5 | 10 | 20 | 50 |
| PCB 118 | 0.2 | 0.5 | 1 | 2 | 5 | 10 | 20 | 50 |
| PCB 123 | 0.2 | 0.5 | 1 | 2 | 5 | 10 | 20 | 50 |
| PCB 126 | 0.2 | 0.5 | 1 | 2 | 5 | 10 | 20 | 50 |
| PCB 156 | 0.2 | 0.5 | 1 | 2 | 5 | 10 | 20 | 50 |
| PCB 157 | 0.2 | 0.5 | 1 | 2 | 5 | 10 | 20 | 50 |
| PCB 167 | 0.2 | 0.5 | 1 | 2 | 5 | 10 | 20 | 50 |
| PCB 169 | 0.2 | 0.5 | 1 | 2 | 5 | 10 | 20 | 50 |
| PCB 189 | 0.2 | 0.5 | 1 | 2 | 5 | 10 | 20 | 50 |
| Internal Standard | ||||||||
| 13C12-PCB 77 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| 13C12-PCB 81 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| 13C12-PCB 105 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| 13C12-PCB 114 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| 13C12-PCB 118 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| 13C12-PCB 123 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| 13C12-PCB 126 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| 13C12-PCB 156 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| 13C12-PCB 157 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| 13C12-PCB 167 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| 13C12-PCB 169 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
| 13C12-PCB 189 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 |
GC/MS/MS calibration and analyte identification
The concentration of the all congeners was determined using nine-point calibration curve(standard solutions CS1 to CS9 as indicated in TABLE 3) for PCDDs/PCDFs and seven-point calibration curve(standard mixture CS1 to CS7 as indicated in TABLE 4) for DL-PCBs. Calculation of relative response factors for native congeners were calculated using following formula:
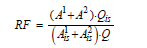
Where,
A1 , A2 = areas of the two native congener diagnostic ions (TABLE 1 and 2)
AIS1 , AIS2 = areas of the two internal standard diagnostic ions (TABLE 1 and TABLE 2).
Q = concentration of corresponding native congener, ng/ml.
QIS = concentration of internal standard relevant to native congener, ng/ml.
Next, the same procedure was applied to calculation of the relative response factor for the internal standards using the appropriate recovery standards using the following formula:
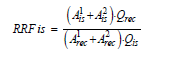
Where,
Arec1 , Arec2 = areas of the two recovery standard diagnostic ions.
Qrec = concentration of corresponding recovery standard, ng/ml.
The limit of detection (LOQ) is not a constant value, it depends by dilutions, weight of the samples, analyte recovery and the sum of all congeners shall be about one fifth of the maximum level [4].
Because S/N (signal/noise) ratio is too small we choose lowest concentration point on a calibration curve approach for determination iLOQ. The LOQ is calculated from the lowest concentration point taking into account the recovery of internal standrds added and the sample intake (Table.5).

TABLE 5. Ion Ratio of native congeners and ISTD recoveries, LOQs values of calculated on the basis
| Congener Name | Ret. TIme | TEF | ISTD Recovery | UB WHO TEQ | MB WHO TEQ |
|---|---|---|---|---|---|
| 2,3,7,8-TCDD | 20.01 | 1 | 1.16 | 0.02 | 0.01 |
| 2,3,7,8-ТCDF | 19.4 | 0.1 | 1.007 | 0.002 | 0.001 |
| 1,2,3,7,8-PCDD | 25.34 | 1 | 0.857 | 0.04 | 0.02 |
| 1,2,3,7,8-PCDF | 23.7 | 0.03 | 1.257 | 0.0012 | 0.0006 |
| 2,3,4,7,8-PCDF | 24.9 | 0.3 | 1.2 | 0.0012 | 0.0006 |
| 1,2,3,4,7,8-HXDF | 29.54 | 0.1 | 0.983 | 0.008 | 0.004 |
| 1,2,3,6,7,8-HXDF | 29.74 | 0.1 | 1.041 | 0.008 | 0.004 |
| 2,3,4,6,7,8-HXDF | 30.6 | 0.1 | 1 | 0.006 | 0.003 |
| 1,2,3,4,7,8-HXDD | 30.88 | 0.1 | 0.646 | 0.0012 | 0.0006 |
| 1,2,3,6,7,8-HXDD | 31 | 0.1 | 0.774 | 0.006 | 0.003 |
| 1,2,3,7,8,9-HXDD | 31.46 | 0.1 | 0.631 | 0.008 | 0.004 |
| 1,2,3,7,8,9-HXDF | 31.9 | 0.1 | 0.749 | 0.008 | 0.004 |
| 1,2,3,4,6,7,8-HPDF | 34.1 | 0.01 | 0.952 | 0.004 | 0.002 |
| 1,2,3,4,6,7,8-HPDD | 35.58 | 0.01 | 0.719 | 0.001 | 0.0005 |
| 1,2,3,4,7,8,9-HPDF | 36.2 | 0.01 | 0.791 | 0.004 | 0.002 |
| OCDD | 39.19 | 0.0003 | 0.719 | 0.00003 | 0.00002 |
| OCDF | 39.38 | 0.0003 | 1.011 | 0.00002 | 0.00001 |
| PCB 77 | 22.52 | 0.0001 | 0.846 | 0.000017 | 0.000008 |
| PCB 81 | 21.86 | 0.0003 | 0.835 | 0.000047 | 0.000002 |
| PCB 123 | 23.89 | 0.00003 | 0.687 | 0.000003 | 0.000001 |
| PCB 118 | 24.14 | 0.00003 | 0.812 | 0.000002 | 0.000001 |
| PCB 114 | 24.81 | 0.00003 | 0.768 | 0.000004 | 0.000002 |
| PCB 105 | 25.85 | 0.00003 | 0.77 | 0.000004 | 0.000002 |
| PCB 126 | 28.24 | 0.1 | 1.041 | 0.02012 | 0.01006 |
| PCB 167 | 29.69 | 0.00003 | 1.011 | 0.000006 | 0.000003 |
| PCB 156 | 31.53 | 0.00003 | 0.932 | 0.000005 | 0.000002 |
| PCB 157 | 31.95 | 0.00003 | 0.978 | 0.000005 | 0.000002 |
| PCB 169 | 35.33 | 0.03 | 0.914 | 0.004578 | 0.002289 |
| PCB 189 | 39.6 | 0.00003 | 1.135 | 0.000006 | 0.000003 |
Results
Calibration
(FIG. 4 - 10) showed mass chromatograms for each reference standard and negative samples for all test analytes. Calibration standards (seven levels for PCDD/Fs and seven levels for DL-PCBs) were analysed for three analytical sequences and demonstrated good RF% RSDs within EU regulations. TABLE 6 shows the data obtained for PCDD/Fs and dioxin-like PCBs. The Relative Response Factors (RRFs) were calculated at each concentration level and the linearity was estimated bades on the RRF and in the Determination Coefficient (R2 ). The Accuracy and Instrumental Limit of Quantification (iLOQ) were assessed for each congener. Accuracy was expressed in terms of bias% and mean squared error and it was measured in standard solutions. The iLOQ was calculated at the lowest calibration point and set by 10 times standard deviation using 10 replicate injections. Finally, precision was expressed as Relative Standard Deviation (RSD) for the calibration curve levels (n = 10 for the lowest calibration point).
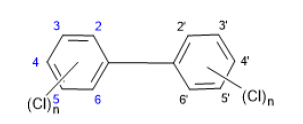
Figure 4: Chromatographic Profiles of TCDD, PCDD and HxDD: where (a) (b) and (c) represent TCDD represent PCDD and HxDD respectively.
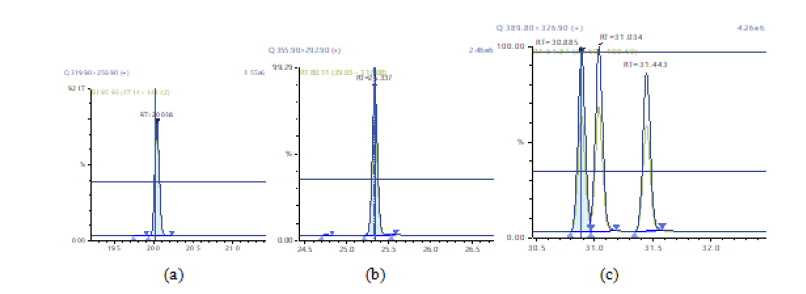
Figure 5: Chromatographic Profiles of HpDD and OCDD: where (a) and (b) represent HpDD OCDD respectively
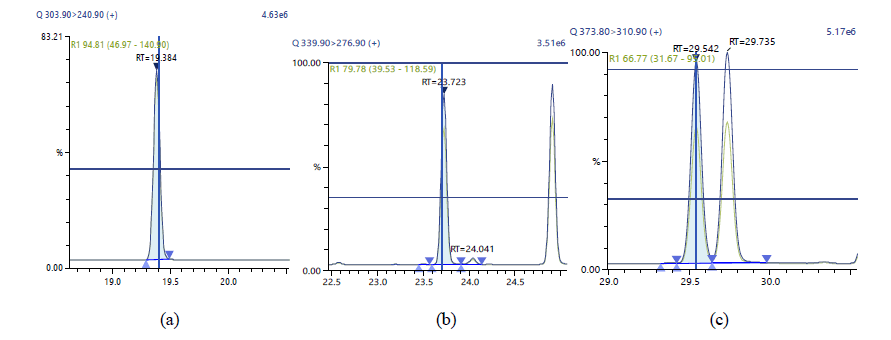
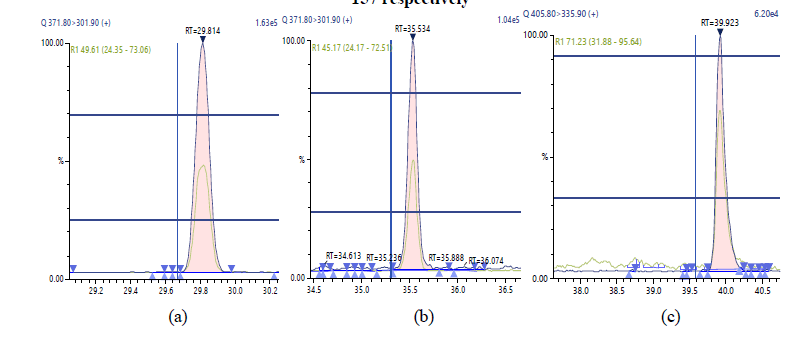
Figure 6: Chromatographic Profiles of TCDF,PCDF and HxDF: where (a), (b) and (c) represent TCDF, PCDFand HxDF respectively
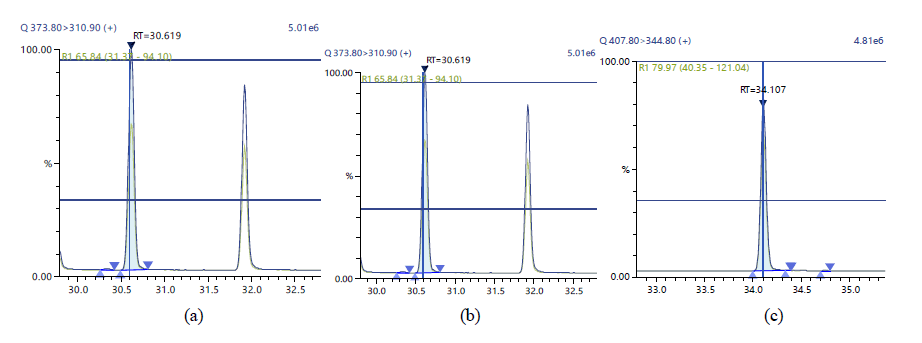
Figure 8: Chromatographic Profiles of DL-PCBs: where (a), (b) and (c) represent PCB 105, PCB 114 and PCB 118
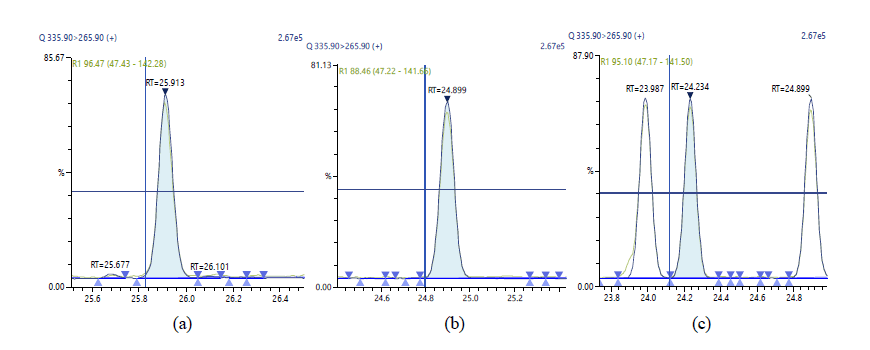
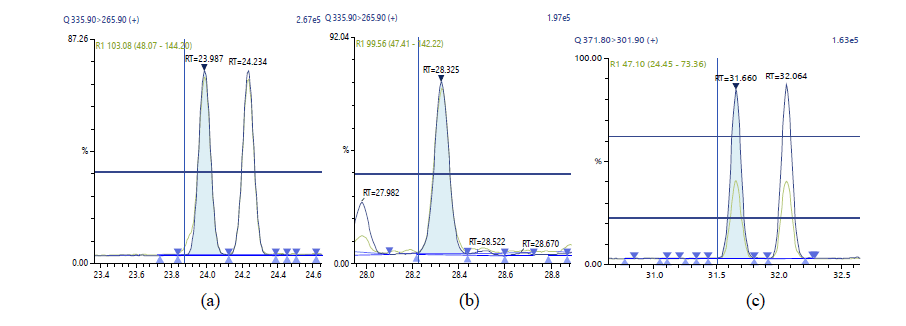
Figure 9: . Chromatographic Profiles of DL-PCBs: where (a), (b) and (c) represent PCB 123, PCB 126 and PCB 156, PCB 157 respectively
Figure 10: Chromatographic Profiles of DL-PCBs: where (a), (b) and (c) represent PCB 167, PCB 169 and PCB 180 respectively
TABLE 6. Calibration data for PCDDs/PCDFs and DL-PCBs congeners
| Peak Name | Retention Time (min) | Coefficient of Determination (R2) | RF RSD (%) | Mean RF (Slope) | Range (pg) |
|---|---|---|---|---|---|
| 2,3,7,8-TCDD | 20.01 | 0.99914 | 2.33 | 2.19 | 0.025-50.00 |
| 2,3,7,8-ТCDF | 19.4 | 0.9994 | 1.85 | 2.21 | 0.025-50.00 |
| 1,2,3,7,8-PCDD | 25.34 | 0.99947 | 1.62 | 1.62 | 0.025-50.00 |
| 1,2,3,7,8-PCDF | 23.7 | 0.99924 | 2.05 | 2.05 | 0.025-50.00 |
| 2,3,4,7,8-PCDF | 24.9 | 0.99953 | 1.77 | 1.77 | 0.025-50.00 |
| 1,2,3,4,7,8-HXDF | 29.54 | 0.99953 | 1.83 | 1.83 | 0.050-100.00 |
| 1,2,3,6,7,8-HXDF | 29.74 | 0.99955 | 1.9 | 1.89 | 0.050-100.00 |
| 2,3,4,6,7,8-HXDF | 30.6 | 0.9996 | 1.51 | 1.51 | 0.050-100.00 |
| 1,2,3,4,7,8-HXDD | 30.88 | 0.99937 | 1.74 | 1.74 | 0.050-100.00 |
| 1,2,3,6,7,8-HXDD | 31 | 0.99947 | 1.64 | 1.64 | 0.050-100.00 |
| 1,2,3,7,8,9-HXDD | 31.46 | 0.99951 | 1.99 | 1.99 | 0.050-100.00 |
| 1,2,3,7,8,9-HXDF | 31.9 | 0.9994 | 1.81 | 1.81 | 0.050-100.00 |
| 1,2,3,4,6,7,8-HPDF | 34.1 | 0.99927 | 1.75 | 1.76 | 0.050-100.00 |
| 1,2,3,4,6,7,8-HPDD | 35.58 | 0.99958 | 2.01 | 2.01 | 0.050-100.00 |
| 1,2,3,4,7,8,9-HPDF | 36.2 | 0.99946 | 1.9 | 1.9 | 0.050-100.00 |
| OCDD | 39.19 | 0.99942 | 1.39 | 1.4 | 0.10-200.00 |
| OCDF | 39.38 | 0.99903 | 1.58 | 1.58 | 0.10-200.00 |
| PCB 77 | 22.52 | 0.99962 | 0.9 | 2.73 | 0.20-50.00 |
| PCB 81 | 21.86 | 0.99941 | 1 | 2.85 | 0.20-50.00 |
| PCB 123 | 23.89 | 0.99951 | 2.85 | 2.93 | 0.20-50.00 |
| PCB 118 | 24.14 | 0.99942 | 0.79 | 3.02 | 0.20-50.00 |
| PCB 114 | 24.81 | 0.99922 | 3.01 | 2.93 | 0.20-50.00 |
| PCB 105 | 25.85 | 0.9992 | 2.93 | 2.96 | 0.20-50.00 |
| PCB 126 | 28.24 | 0.99929 | 0.86 | 2.88 | 0.20-50.00 |
| PCB 167 | 29.69 | 0.99937 | 0.69 | 2.87 | 0.20-50.00 |
| PCB 156 | 31.53 | 0.99926 | 0.73 | 3 | 0.20-50.00 |
| PCB 157 | 31.95 | 0.99947 | 0.69 | 2.93 | 0.20-50.00 |
| PCB 169 | 35.33 | 0.99932 | 0.82 | 2.96 | 0.20-50.00 |
| PCB 189 | 39.6 | 0.99938 | 0.75 | 2.85 | 0.20-50.00 |
The lowest acceptable calibration point was determined according two criteria. First, the calculated RSDs of the lowest level for all congeners must be ≤ 15%. Second, the relative difference between the RRF average obtained for all levels and the RRF average obtained for the lowest level must be ≤ 30%, according to the regulation. As can be observed in TABLE 6, this criteria was met and linearity was acceptable within the calibration range. At this point, the lowest calibration level was used to determine the Iloq.
Reaching the level of interest
Each congener has different strength of toxicity, and is expressed as Toxic Equivalent Factor (TEF). The TEF value of 2,3,7,8- TCDD has values of 1, which is the most toxic congener.
All the regulated target compounds include the most toxic congeners of PCDD/Fs and DL-PCBs which have a toxic equivalent factor assigned by the World Health Organization (WHO) [5-6].
The maximum acceptable level for the PCDDs/PCDFs and DL-PCBs in foods and feeds are prescribed by their Toxic Equivalents (TEQ). The TEQ was calculated by multiplying the concentration of each compound by the TEF, and then calculating the total TEQ for all congeners.
In this study, two types of vegetable oils (rapeseed and sunflower oil), and sunflower meal pellets were analysed using GC/MS/MS. The LOQ of each individual congener was calculated from the lowest concentration point (CS1) taking account the recovery of internal standards (60%-120%) and ion abundance.
Quantification of PCDD/PCDFs and DL-PCBs in sample extracts
Following successful validation of the method, the corresponding congeners were quantified in the sample extracts. Excellent chromatographic separation with minimal matrix interference was observed for all labelled congeners in all sample extracts analysed.
The Ion Ratio (IR) abundance for selected transitions of each of PCDD/PCDFs and DL-PCBs congeners was measured in each of the samples analysed and the values were compared with measured ion ratio values (average from the calibration standards CS1- CS9).The results of this study show that all the IR for the analysed compounds were within the 15% tolerance, meeting the EU criteria for dioxin confirmation [7-10].
Discussion
The advantages of the new approach are obvious. Combination of short extraction time, use of small solvent volume and high performance clean-up strategy results in short delivery time and high quality chromatograms which are easy to process. The extraction of maize, wheat and sunflower meals starts with sample intake of 10 gm and the required volume of organic solvent is 70 ml. The total extraction time required is approximately 1 hour. At the end of extraction time, the solvent is concentrated down to near dryness the final extract fat/oil and put directly on clean-up column. After purification the obtained two fractions are evaporated to 20 µl in GC vial equipped with insert and put directly in an auto-sampler. In the case of vegetable oil, the samples were subjected to acid digestion before multilayer column cleanup and the remaining procedure as same as mentioned above(FIG.11-18)..
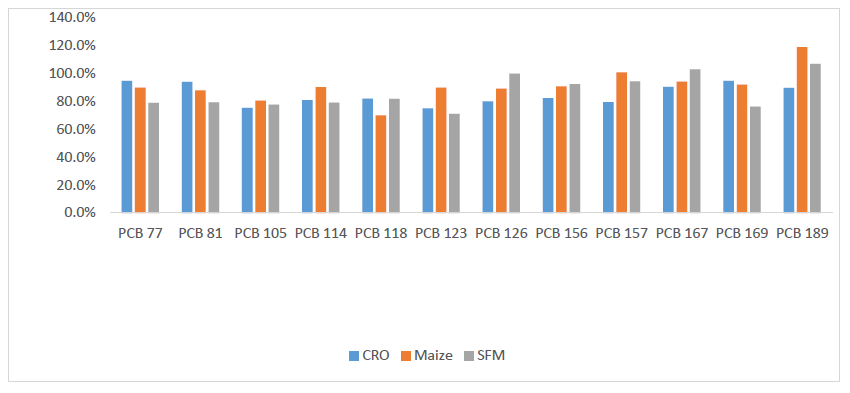
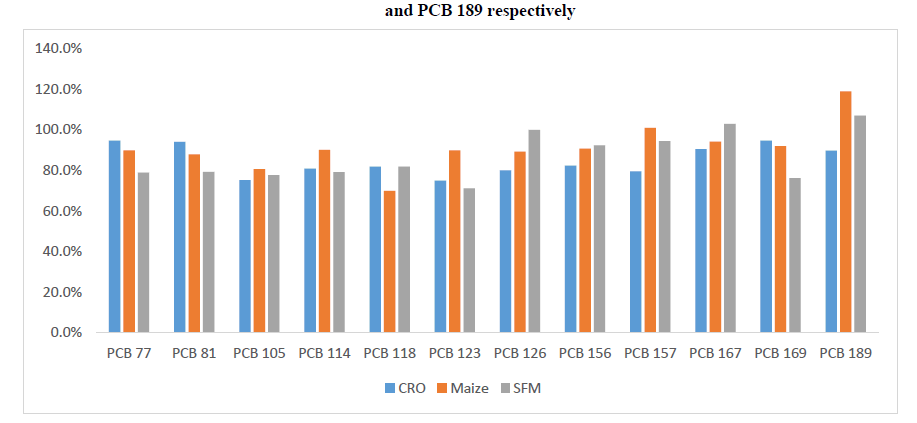
Figure 11: PCDD/PCDFs data for Sunflower Feed Meal (SFM), crude sunflower oil (CSO) and crude rapeseed oil (CRO)
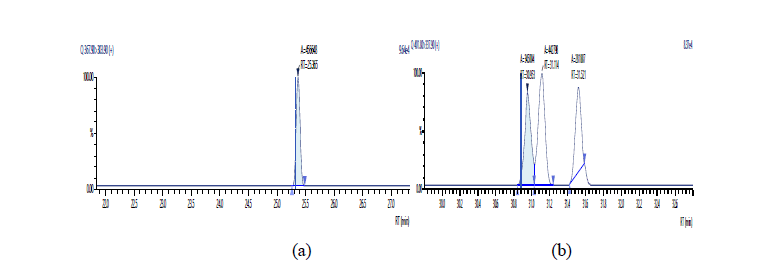
Figure 12: Chromatographic Profiles of PCDD/PCDFs in maize extraxt: where (a) and (b) represent TCDD and HxDD respectively
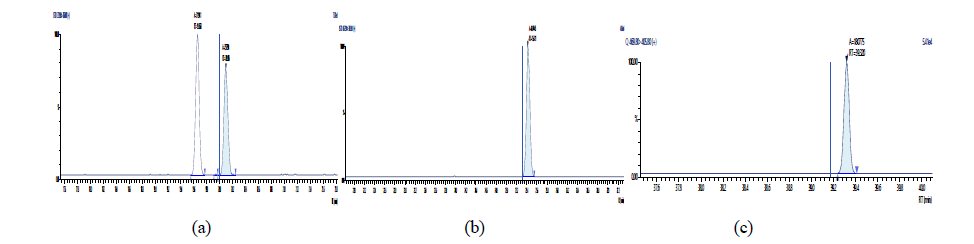
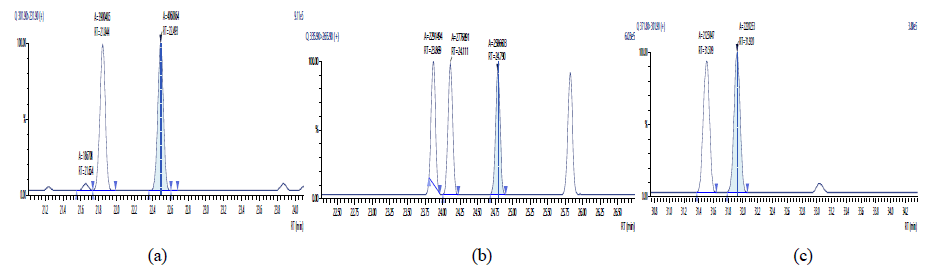
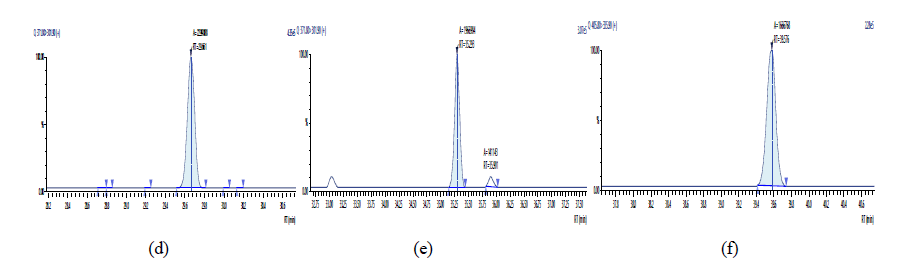
Figure 13: Chromatographic Profiles of PCDD/PCDFs in crude sunflower oil: where (a), (b) and (c) represent the TCDD, PCDD and OCDD respectively.
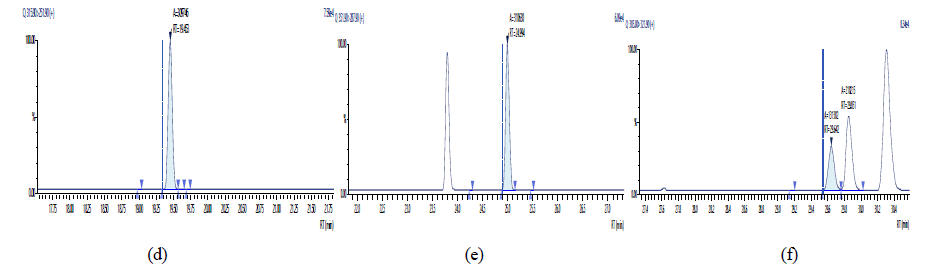
Figure 14: Chromatographic Profiles of PCDD/PCDFs in crude sunflower oil: where (d), (e) and (f) represent TCDF, PCDF and HxDF respectively
Figure 15: Chromatographic Profiles of PCDD/PCDFs in crude sunflower oil: where (g) and (h) represent HpDF and OCDF respectively.
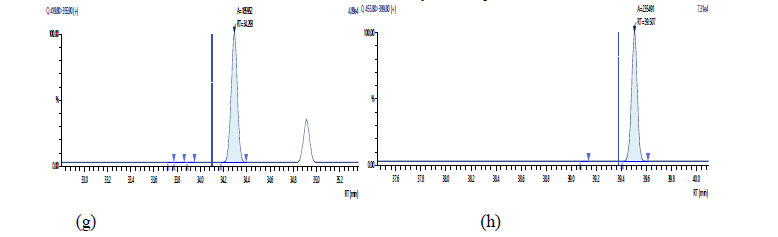
Figure 16: Chromatographic Profiles of DL-PCBs in crude rapeseed oil: where (a) represent PCB 77, PCB 81 (b) PCB 105,114,118,123 and (c) PCB 156,157
Figure 17: Chromatographic Profiles of DL-PCBs in crude rapeseed oil: where (d), (e) and (f) represent PCB 167, PCB 169
Figure 18: DL-PCBs data for Sunflower Feed Meal (SFM), Crude Sunflower Oil (CSO) and Crude Rapeseed Oil (CRO
Conclusions
A GC/triple quadrupole MS/MS method has been developed and fully validated in accordance with criteria in EU Regulation 709/2014 that allows the use of GC/triple quadrupole MS/MS as a confirmatory method for official control of PCDDs, PCDFs and DL-PCBs in animal feedstuffs and vegetable oils. This method meets the requirements of the regulation, and can achieve similar performance to GC/HRMS. The clear and confident detection of these congeners at very low levels, even within typically complex food and feed matrices, illustrates the high sensitivity of the method, as well as its high analytical value considering the extremely high toxicity of many dioxins. Finally, the stability of the target ion ratios across the entire set of analyses-including a range of PCDD/PDFs and DL-PCBs concentrations and potential matrix effects-supports the utility and reliability of this method for high confidence analyses to meet current regulatory demands.
References
- Method 1613. Tetra-through octa-chlorinated dioxins and furans by isotope dilution HRGC/HRMS. Natl serv cent environpubl. 1993.
[Google Scholar] [Crossref]
- Method 1668. Chlorinated biphenyl congeners in water, soil, sediment, and tissue by HRGC/HRMS. 2010
[Google Scholar] [Crossref]
- Takakura M, Lehardy T, Marchard P. Analysis of dioxins in foods and feeds using GC-MS/MS. 2018.
-
[Google Scholar] [Crossref]
- Ellison SLR, Rosslein M, Williams A. Quantifying uncertainty in analytical measurement. Eurachem quam. 2000;1:120.
[Google Scholar] [Crossref]
- V M, Birnbaum LS, Denison M, et al. The 2005 World Health Organization reevaluation of human and mammalian toxic equivalency factors for dioxins and dioxin-like compounds. Toxicol sci. 2006;93(2):223–41.
- Furst P. Contribution of dioxin-like PCB to total toxic equivalents of dairy products. Organohalogen compd. 2001;51:279-82.
[Google Scholar] [Crossref]
- Setting maximum levels for certain contaminants in foodstuffs as regards dioxins and dioxin-like PCBs. European Commission (EC). Off J. 2006;34-8.
[Google Scholar] [Crossref]
- Li Z, Sun Y, Dong F, et al. Discrimination of heating and frying vegetable oils based on UPLC/Q-TOF MSMS and chemometrics. Food Res Int. 2021;140:109874.
- Chen L, Liu Y, Aipeng HU, et al. Effect of soybean oil deodorization and distillate trapping process on the quality of products. J Henan Univ Technol. 2018:39:43–50.
- Lucci P, Bertoz V, Pacetti D, et al. Effect of the Refining Process on Total Hydroxytyrosol, Tyrosol, and Tocopherol Contents of Olive Oil. Foods. 2020;9:292.