Research
, Volume: 14( 6)Primary Evaluation of the Cold Shock Protein Existing in the Strain Lactobacillus Plantarum Ul497 and the StrainâÂÂs Survival after Temperature Downshock
- *Correspondence:
- Uyen Quynh Nguyen , Laboratory of Enzyme-Protein, Institute of Microbiology and Biotechnology, Vietnam National University, Hanoi, Vietnam, E-Mail: uyennq@vnu.edu.vn
Received: October 03, 2018; Accepted: October 30, 2018; Published: November 05, 2018
Citation: Nguyen UQ, Vo LTT. Primary Evaluation of the Cold Shock Protein Existing in the Strain Lactobacillus Plantarum Ul497 and the Strain?s Survival after Temperature Downshock. Biotechnol Ind J. 2018;14(6):177.
Abstract
Lactic Acid Bacteria (LAB) have been approved by FDA as GRAS (Generally Recognized as Safe). Among LAB, Lactobacillus plantarum along with other species in the genus Lactobacillus shows its roles in many fields, especially in the food industry. In this study, the deduced amino acids of cold shock protein (Csp) in the strain Lactobacillus plantarum UL497 were compared with those of the reported cold shock proteins, including CspL, CspC and CspP of Lactobacillus plantarum NC8. Then, after freezing, the survival of the bacteria with temperature downshift from 30°C to 8°C for 2, 5 and 8 hours at the growing log phase was determined. In particular, after freezing, the survival percentage of the bacteria with temperature downshift from 30°C to 8°C at the growing log phase increased in comparison with that of bacteria without temperature downshift. Also, based on these survival percentages, the effects of duration of cold shock on the bacteria’s survival were primitively evaluated.
References
Best Betting Sites in Turkey Blog | Mexican Food Recipes Casino Sites in Argentina Best Betting Sites in Canada Blog | Turkish Food Recipes Fan Tokens Info Blog | Vietnamese Food Recipes Engine Technologies Articles Blog | Yacht Charter in UAE All About Digital Coins Blog | Yacht Charter in Balkans Casino Sites in Bangladesh Blog | Yacht Charter in Australia Mortgage info in New York Blog | Yacht Charter in China Best Betting Sites in Nigeria Blog | Yacht Charter in Spain Best Betting Sites in United Arab Emirates Blog | Yacht Charter in France Places to Visit in ChinaKeywords
Amino acids; Cold shock proteins; Lactic Acid Bacteria (LAB); Lactobacillus plantarum
Introduction
Lactic Acid Bacteria (LAB) have been approved by FDA as GRAS (Generally Recognized as Safe). LAB is capable of producing variable bioactive compounds, including bacteriocins, organic acids (like lactic acid and acetic acid), exopolysaccharides and other valuable substances which play important roles not only in living organisms but also in many fields in our life such as in medicine and the food industry [1-3]. In the latter, Lactobacillus plantarum is found in different kinds of fermented foods playing the role of starter cultures or preservatives with a natural origin. Therefore, L. plantarum has been the object of many studies.
Csp(s) are family of cold shock proteins that are found in most organisms, including different species of LAB related to foods. The molecular weights of these proteins are around 7 kDa and their amino acid sequences display a high similarity to one another. The functions of these proteins have been investigated in many studies and one of them is the contribution of the proteins in the adaptation process of bacteria to new environmental conditions, especially in temperature downshift [4,5]. For Lactobacillus plantarum, three csp proteins, including cspC, cspL, and cspP are reported. The cspC protein is 73% and 74% identical to cspL and cspP, respectively, whereas cspL and cspP are 88% identical to each other [6].
In this study, based on the nucleotide sequences the deduced amino acids of the Csp protein existing in the strain Lactobacillus plantarum UL497 were obtained. These sequences were aligned with those of the reported CspC, CspL and CspP in L. plantarum NC8. Then, after freezing, the survival percentages of the bacteria with temperature downshift from 30°C to 8°C for 2, 5 and 8 hours at the growing log phase were determined. Based on the values of these survival percentages, the effects of duration of cold shock on the bacteria’s survival were primitively evaluated.
Materials and Methods
Bacterial strains
The strain UL497 was isolated from the fermented tofu “Chao” of Hue province (Vietnam). Based on the 16S rDNA gene sequencing and sugar fermentation pattern analysis by using API CH50 (bioMérieux SA, USA), the strain was identified as L. plantarum. The strain was stored at -80°C in MRS medium (Oxoid, Basing-stoke, United Kingdom) with 15% glycerol, and before use, it was propagated in MRS medium at 30°C for 24-48 hours. The indicator strain used in this study was Lactobacillus sakei subsp. sakei JCM1157.
All chemicals and media, including MRS and LB media were purchased from Promega (USA), Sigma Aldrich (USA) and LAB (Neogen company, USA).
PCR and cloning
DNA genome extraction and competent cells of Escherichia coli DH5α were performed following the protocol of Sambrook J. et al [7]. The DNA genome of the strains was used as the templates for the amplification of Csp genes by using the primers (Csp1a: 5’ GGTTTAATGTAGACAARGGNTTYGGNTTYAT 3’ and Csp1b: 5’ TAGTAGGTACCRTTNGCNGCYTGNGGNCC 3’) with the expected size of 178 bp of PCR products [8,9]. The reaction mixture (50 μL) contained 5 μL of 10X master mix buffer, 50 pmol of each primer, 2.5 U of Taq DNA polymerase, and 1 μL of template DNA. Thermocycles of PCR included 5 minutes of heat shock at 95°C, followed by 30 cycles of 95°C for 30 seconds, 55°C for 30 seconds, and 72°C for 30 minutes and a final extension at 72°C for 10 minutes. The PCR product was checked by electrophoresis on 1% agarose gel, purified, cloned into pGEM-T vector (Promega) and then transformed into the competent cells of Escherichia coli DH5α. After that, the sample was sent to the 1st Base company (Singapore) for sequencing. The Clustal Omega, ExPASy tools were used to analyze the nucleotide and amino acid sequences of the Csp gene and Csp protein existing in the strain.
Survival assessment
The strain was grown in MRS media at 30°C for 2.5-4 hours until the OD600 of the culture reached 0.3-0.5. The cultures were divided into several samples: one sample was immediately frozen at -20°C, and the other were cold shocked from 30°C to 8°C for 2, 5 and 8 hours, respectively before they were frozen at -20°C [6,8-10]. For each sample, before and after freezing for 24 hours at -20°C, the Colony Forming Units (CFU) obtained by serially diluting in sterile water were counted to give a percentage of viable cells for each sample. All experiments were done in triplicate. The T-test was used for statistics.
Results and Discussion
Cold shock protein existing in the strain L. plantarum UL497
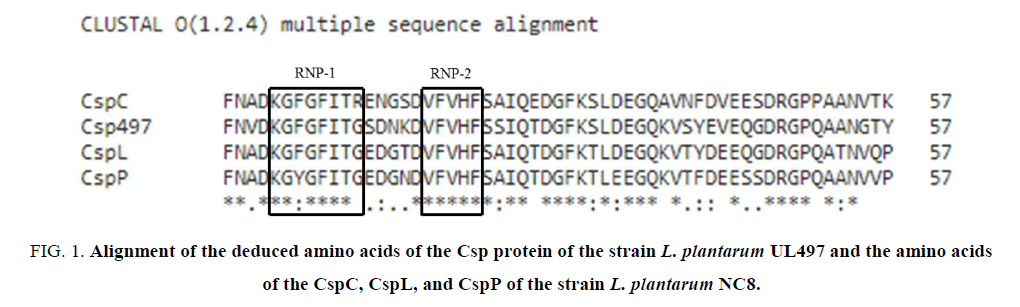
The PCR product of the csp gene amplification of the strain L. plantarum UL47 was checked, cleaned and then cloned into a pGEM-T vector (Promega) following the instruction of the manufacturer. After getting the nucleotide sequences (the result not shown here) the deduced amino acids of the Csp protein existing in the strain L. plantarum UL497 were aligned with those of the reported CspC, CspL and CspP in the strain L. plantarum NC8 (FIG. 1).
Figure 1: Alignment of the deduced amino acids of the Csp protein of the strain L. plantarum UL497 and the amino acids of the CspC, CspL, and CspP of the strain L. plantarum NC8.
The deduced amino acids from the nucleotide sequences obtained after sequencing showed that the Csp protein of the strain L. plantarum UL497 contained 2 conserved RNA binding motifs (RNP-1 and RNP-2). The percentages of the deduced amino acids of the Csp protein of the strain UL497 that were identical and similar to those of the reported Csp proteins, including CspC, CspL, and CspP of L. plantarum NC8, were shown in TABLE 1.
| Csp proteins | Percent of the deduced amino acids of the Csp protein of the strain UL497 that was identical to others | Percent of the deduced amino acids of the Csp protein of the strain UL497 that was similar to others |
|---|---|---|
| CspL | 77.19% | 85.96% |
| CspP | 70.18% | 84.21% |
| CspC | 70.18% | 80.70% |
Table 1: The percentages of the deduced amino acids of the Csp protein of the strain UL497 that was identical and similar to those of the CspC, CspL and CspP of the strain L. plantarum NC8.
The result showed that among 3 Csp proteins that already reported in the L. plantarum NC8, Csp protein existing in the strain L. plantarum UL497 was closest to the CspL. The lowest percents of the deduced amino acids of the Csp protein of the strain UL497 that were identical and similar to those of CspC in the L. plantarum NC8 were 70.18% and 80.70%, respectively.
The survival of bacteria after temperature downshift
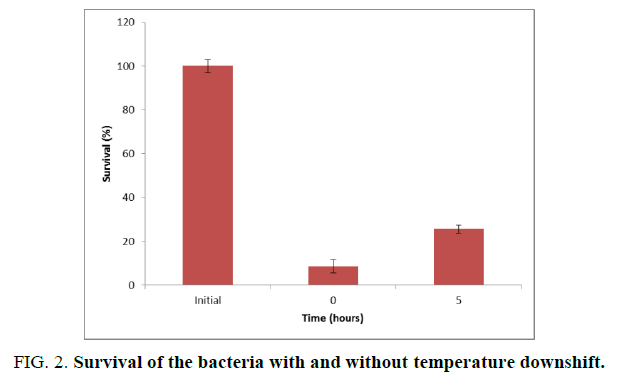
Based on the analysis of the deduced amino acids of the Csp protein existing in the strain L. plantarum UL497, at first, the survival of the bacteria after temperature downshift (from 30°C to 8°C) for 5 hours at its growing log phase was evaluated (FIG. 2).
The result showed that the survival of the bacteria immediately frozen at -20°C drastically decreased (remained at 8.63%), but it increased when the bacteria were cold shocked (from 30°C to 8°C) at the growing log phase for 5 hours (obtained 25.61%).
Effects of duration of cold shock on the survival of the bacteria
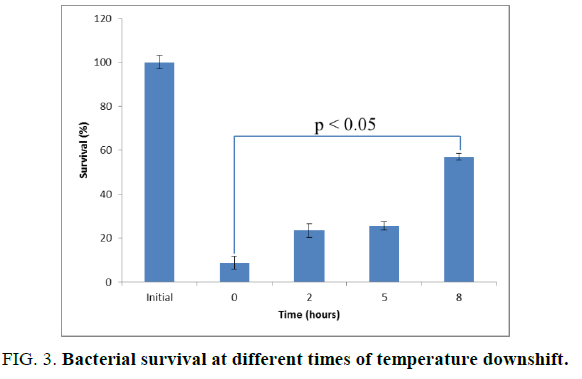
To assess whether the duration of cold shock affects the survival percentages of the bacteria, the cultures were cold shocked for 2, 5 and 8 hours, and then their survivals were determined after freezing at -20°C. The results were plotted in FIG. 3.
After freezing, all of the bacterial samples with cold shock for 2, 5 and 8 hours increased their survival percentages compared to the bacteria without cold shock. Notably, the survival of the bacteria cold shocked for 2, 5 and 8 hours was 2.7, 2.9 and 6.5 times higher respectively than that of non-cold shocked bacteria. In addition, the survival of the bacteria cold shocked for 8 hours showed statistically significant differences from that of non-cold shocked ones with the p-value of 0.002 determined by T-test.
Discussion
Lactobacillus plantarum is widely applied in many variable fermented foods with the role of starter cultures or preservatives. In the food industry, the bacteria are exposed to processing and storage conditions, including temperature downshift. Therefore in this field, the bacteria’s ability to survive in different conditions has been of an important issue. In our study, in the strain L. plantarum UL497, Csp protein which is thought to play an important role in bacterial adaptation to cold stress was analyzed, and then after freezing the survival of the bacteria with temperature downshift from 30°C to 8°C for 2, 5 and 8 hours at the growing log phase was primitively evaluated.
The deduced amino acids of the Csp protein in the strain UL497 showed that the protein possessed typical characteristics of the proteins belonging to the Csp protein family. In particular, it contained 2 conserved RNA binding motifs (RNP-1 and RNP-2) via which the protein fulfills its functions during the adaption process [4,5]. In addition, the identity and similarity of the deduced amino acids of this protein with those of all three Csp proteins in L. plantarum NC8 were high. In particular, the highest identity and similarity were observed between the deduced amino acids of the Csp protein of the strains L. plantarum UL497 and CspL with the values of 77.19% and 85.96%, respectively [6].
Many publications showed the roles of Csp proteins, especially in cold adaptation of bacterial cells. However, the specific actions and the mechanism via which Csp(s) interacts to fulfill their functions at both normal and stress growing conditions, including cold shock, have not been clear yet. mRNA of all three csp genes decreased at the stationary growing phase of L. plantarum [10], therefore the role of the Csp protein existing in the strain L. plantarum UL497 was tested when the strain’s temperature was downshifted from 30°C to 8°C for 2, 5 and 8 hours at its growing log phase. Our result showed that after freezing the survival percentages of the bacteria with temperature downshift for all time-tested here increased. Interestingly, only the survival of the bacteria cold shocked for 8 hours showed significant differences compared to that of non-cold shocked ones (p<0.05) and there were no significant differences observed for the bacteria’s survival in the rest of samples. It was already known that the level of cspL mRNA increased up to 3-8 times after 3 hours at 8°C, and these levels of cspC and cspP genes remained unchanged after the temperature downshift [6].
Our result here showed that the Csp protein of the strain L. plantarum UL497 displayed the highest identity (77.19%) and similarity (85.96%) in amino acids sequences with those of CspL in L. plantarum NC8. In our previous study, Csp of L. plantarum UL485 was identified as CspC with 91.23% and 94.74% identity and similarity, respectively, in the amino acid sequences of CspC in L. plantarum NC8. Moreover, no significant differences of the survival percentages were observed in the bacteria with and without temperature downshift even though the experimental conditions were the same (cold shock from 30°C to 8°C for 2, 5 and 8 hours at the growing log phase) [11]. Therefore, the reasons to explain survival differences of the bacteria with and without temperature downshift in various durations are still a question that needs further investigation in order to increase the survival of L. plantarum.
Conflicts of Interest
The authors declare no conflicts of interest.
Acknowledgment
This research is supported by “Evaluation of typical Lactic Acid Bacteria applicable in foods, pharmaceuticals and animal feeds” grant of the Ministry of Science and Technology.
References
- De Vuyst L, Leroy F. Bacteriocins from lactic acid bacteria: production, purification, and food applications. J Mol Microbiol Biotechnol. 2007;13(4):194-99.
- Khalid K. An overview of lactic acid bacteria. Int J Biosci. 2011;1(3):1-3.
- Cotter PD, Hill C, Ross RP. Food microbiology: bacteriocins: developing innate immunity for food. Nat Rev Microbiol. 2005;3(10):777.
- Barria C, Malecki M, Arraiano CM. Bact. Adapt Cold Microbiol. 2013;159(12):2437-43.
- Wouters JA, Rombouts FM, Kuipers OP, et al. The role of cold-shock proteins in low-temperature adaptation of food-related bacteria. System Appl Microbiol. 2000;23(2):165-73.
- Derzelle S, Hallet B, Francis KP, et al. Changes in cspL, cspP, and cspCmRNA abundance as a function of cold shock and growth phase in Lactobacillus plantarum. J Bacteriol. 2000;182(18):5105-13.
- Sambrook J, Fritsch EF, Maniatis T. Molecular cloning: a laboratory manual. Cold spring harbor laboratory press; 2001.
- Kim WS, Dunn NW. Identification of a cold shock gene in lactic acid bacteria and the effect of cold shock on cryotolerance. Curr Microbiol. 1997;35(1):59-63.
- Kim WS, Khunajakr N, Ren J, et al. Conservation of the major cold shock protein in lactic acid bacteria. Curr Microbiol. 1998;37(5):333-6.
- Derzelle S, Hallet B, Ferain T, et al. Improved adaptation to cold-shock, stationary-phase, and freezing stresses in Lactobacillus plantarum overproducing cold-shock proteins. Appl Environ Microbiol. 2003;69(7):4285-90.
- Nguyen UQ, Dong AHT, Nguyen LMT. Some characteristics of the bacteriocin and cold shock protein of the strain Lactobacillus plantarum Ul485 isolated form chao of hue province in vietnam. Biotechnol Ind J. 2018;14(5): 171-78.