Original Article
, Volume: 12( 12)PMTâÃâ¬ÃâProtein Mining Tool
- *Correspondence:
- Madhu B , MS Ramaiah Institute of Technology, Bangalore-54, Karnataka, India, Tel: 080-2360-8769; E-mail: madhoobhan@yahoo.co.in
Received: August 24, 2016; Accepted: September 08, 2016; Published: September 15, 2016
Citation: Madhu B, Anusha KV. PMT – Protein Mining Tool. Biotechnol Ind J. 2016;12(12):110.
Abstract
Proteins play an important role in human body. Composed of amino acids these molecules catalyze reactions in our bodies, transport oxygen, strengthen our immune system and transmit signals from cell to cell. Each protein has its own characteristics and few parameters that the user needs to calculate for understanding the structure, functionality, nature, location of the protein and many other details. Proteome which is the entire set of proteins in a cell differs from cell to cell. Also Proteome of diseased cells is different from proteome of a normal cell. To understand the changes that occur as a result of disease, it is important to understand the normal proteome of a cell. This understanding can result in knowing the molecular basis for the disease, which in turn can help to develop treatment strategies. In this paper we present a tool used for computation of few properties of the proteins which can help the users in the field of proteomics or similar fields so that they can process many huge chunks of uncharacterized proteins and get familiar with them. In this way there are possibilities to identify drugs to cure diseases with respect to proteins.
Keywords
Proteomics; Protein sequences; Motifs; Disordered proteins; Amino acids; Molecular weight; Iso-electric point; Classification; Hydrophobicity; Hydrophilicit
Introduction
Proteomics is to know about composition, structure, function, and interactions of the proteins to determine the activities of each living cell. Known as building blocks of life, proteins are large sized bio-molecules, consisting of chains of amino acids. Proteins are responsible for catalization, body metabolism, replication of DNA, and transportation of molecules. It is majorly the sequence of amino acids which distinguishes one protein from another. Classification of proteins can be done on various parameters like shape, function and structure [1]. Based on shape proteins can be classified into globular and fibrous. Globular proteins are generally soluble in water. Fibrous proteins are commonly found in animals, and are not soluble in water. Based on composition and solubility, proteins are classified as simple proteins or complex proteins. Simple proteins are made of only one type of amino acid, and liberate these amino acids on decomposition with acids. Complex proteins are proteins that are made of amino acids and other organic compounds. The non-amino acid group is termed as prosthetic group. Based on their metabolic function proteins are grouped as enzyme proteins, structural proteins, transport proteins, nutrient and storage proteins, motile proteins, defense proteins, regulatory proteins and toxic proteins. Enzyme proteins are highly specialized proteins with catalytic activity. Structural proteins help in strengthening biological structures. Transport proteins aid in transporting ions or molecules in the body. Nutrient and storage proteins provide nutrition to growing embryos and store ions. Motile proteins function in the contractile system. Defense proteins defend against other organisms. Regulatory proteins regulate cellular or metabolic activities. Toxic proteins hydrolyze or degrade enzymes.
Based on the structure [2], we identify proteins with primary structure, secondary structure, tertiary structure and quaternary structure. Primary structure of protein is the linear sequence of amino acids. This linear sequence of amino acids makes up the polypeptide chain. The secondary structure is a structure of polypeptide chain that cannot be folded and assumes helical shape. Tertiary structure of proteins is the three-dimensional structure formed by the bending and twisting of the polypeptide chain. Some proteins contain more than one polypeptide chains; this kind of polypeptide chains refers to the quaternary structure.
"Protein mining tool" is a product that provides user an interface to handle many protein sequences present in a Fasta file [3]. With this tool the user processes the sequences to compute the molecular weight, iso-electric point and classifies them into hydrophobic and hydrophilic proteins. The tool scans protein sequences for the presence of motifs [4], present in the file. The tool is also used to identify the presence of intrinsically disordered proteins (IDPs) [5]. The identification of motifs and IDPs is further used to infer the functionality of protein sequences. This is achieved through graphical presentation of various characteristics and properties of protein sequences. Overall the tool can help the users to analyze a bulk of proteins sequences and this information can be further used in the field of bioinformatics and proteomics.
The objective of this tool is to handle a large number of sequences at a time to optimize the work of the user where they need not type the sequences individually.
Classification
Molecular mass or molecular weight and iso-electric point of a protein can help the user understand the particular protein’s behavior. Further classifying them into hydrophobic and hydrophilic proteins i.e., to know if they are soluble or insoluble in water will provide a brief idea about the functionality of the proteins [6]. The extent to which proteins attract water is termed as hydrophilicity, and the extent to which proteins repel water is termed as hydrophobicity. Molecular mass is the mass of a molecule computed as sum of the masses of each constituent atom multiplied by the number of atoms of that element. Proteins are separated by their iso-electric point (pI), which is the pH at which the amino acid is neutral. The iso-electric point is determined by seven charged amino acids: glutamate, aspartate, cysteine, tyrosine, lysine and arginine. Nowadays iso-electric points can be obtained by computer programs which can later be identified experimentally.
Motif Search
A protein motif possesses a three-dimensional structure consisting of a particular sequence of amino acids which is often associated with a particular function. Motifs can determine which proteins or protein sequences belong to a given protein family. A simple motif is thus some pattern which is strictly shared by all members of the group, e.g., WTRXEKXXY (where X stands for any amino acid). Protein sequence motifs are signatures of protein families and can often be used as tools for the prediction of protein function. A sequence motif or amino-acid sequence is a pattern that is widespread and has a biological significance. Helix-Loop-Helix is an example of motif which binds to DNA. Domains [7] are discrete portions of proteins that fold independently to have their own function. Scientists have stored and classified domains and motifs in a number of databases. This has helped to analyze proteins for the presence of motifs and domains.
IDP Identification
An IDP is a protein that does not possess ordered three dimensional structure. A proteins function depends on a fixed three-dimensional structure. It so happens that IDPs can adopt an ordered three-dimensional structure after binding to some other macromolecules. Overall, IDPs tend to behave differently in terms of function, structure, sequence, interactions, evolution and regulation [8]. When there is a high hydrophobicity level and low iso-electric point then there is repulsion causing disturbances in the bonding of the protein chains. This is one of the main reasons for the cause of IDPs. IDPs are highly present in many human diseases, including neuro-degeneration and other protein dysfunction maladies and, as such can give scope to new drugs.
Experimental Setup
The existing systems classify the proteins with respect to the similarity with other proteins. Although these systems also calculate the molecular weight, iso-electric point and identifies if any motif is present but it is done for a single sequence. Our proposed tool accepts the protein sequences from a Fasta file [3]. In this file there exist many protein sequences which are uncharacterized and placed one after the other following the Fasta format. These sequences are accepted by the tool and processed for calculating their molecular weight and iso-electric point, identifying if there exists any specific motif. It also helps in analyzing the parameters of the sequences in the file and check if there are any intrinsically disordered proteins. A Fasta file is a text-file used for representing protein sequences or amino acids using single-letter codes. The format of this file allows for sequence names and comments to precede the actual sequences. The format has become a standard in analyzing bioinformatics data. The comment line is distinguished from the sequence data by the symbol ">" in the first column. Lines of text should be shorter than 80 characters in length Table 1.
| Description | Length of the sequence | Molecular weight in KD | Isoelectric point KpI | Nature | Nature % |
|---|---|---|---|---|---|
| >sp|Q6GZX4|001R_FRG3G Putative transcription factor 001R OS=Frog virus 3 (isolate Goorha) GN=FV3-001R PE=4 SV=1 | 256 | 34.33 | 1.61 | Hydrophobic | 42% |
| >sp|Q6GZX3|002L_FRG3G Uncharacterized protein 002L OS=Frog virus 3 (isolate Goorha) GN=FV3-002L PE=4 SV=1 | 320 | 40.40 | 1.92 | Hydrophobic | 39% |
| >sp|Q197F8|002R_IIV3 Uncharacterized protein 002R OS=Invertebrate iridescent virus 3 GN=IIV3-002R PE=4 SV=1 | 458 | 62.16 | 2.61 | Hydrophilic | 37% |
| >sp|Q197F7|003L_IIV3 Uncharacterized protein 003L OS=Invertebrate iridescent virus 3 GN=IIV3-003L PE=4 SV=1 | 156 | 19.84 | 0.95 | Hydrophobic | 36% |
| >sp|Q6GZX2|003R_FRG3G Uncharacterized protein 3R OS=Frog virus 3 (isolate Goorha) GN=FV3-003R PE=3 SV=1 | 438 | 56.17 | 2.62 | Hydrophobic | 47% |
| >sp|Q6GZX1|004R_FRG3G Uncharacterized protein 004R OS=Frog virus 3 (isolate Goorha) GN=FV3-004R PE=4 SV=1 | 60 | 7.58 | 0.36 | Hydrophobic | 43% |
| >sp|Q197F5|005L_IIV3 Uncharacterized protein 005L OS=Invertebrate iridescent virus 3 GN=IIV3-005L PE=3 SV=1 | 217 | 27.76 | 1.27 | Hydrophobic | 39% |
| >sp|Q6GZX0|005R_FRG3G Uncharacterized protein 005R OS=Frog virus 3 (isolate Goorha) GN=FV3-005R PE=4 SV=1 | 204 | 27.08 | 1.18 | Hydrophobic | 39% |
| >sp|Q91G88|006L_IIV6 Putative KilA-N domain-containing protein 006L OS=Invertebrate iridescent virus 6 GN=IIV6-006L PE=3 SV=1 | 352 | 47.99 | 2.12 | Hydrophilic | 39% |
| >sp|Q6GZW9|006R_FRG3G Uncharacterized protein 006R OS=Frog virus 3 (isolate Goorha) GN=FV3-006R PE=4 SV=1 | 75 | 10.18 | 0.44 | Hydrophobic | 41% |
| >sp|Q6GZW8|007R_FRG3G Uncharacterized protein 007R OS=Frog virus 3 (isolate Goorha) GN=FV3-007R PE=4 SV=1 | 128 | 16.00 | 0.83 | Hydrophobic | 41% |
| >sp|Q197F3|007R_IIV3 Uncharacterized protein 007R OS=Invertebrate iridescent virus 3 GN=IIV3-007R PE=4 SV=1 | 447 | 60.07 | 2.72 | Hydrophilic | 39% |
| >sp|Q197F2|008L_IIV3 Uncharacterized protein 008L OS=Invertebrate iridescent virus 3 GN=IIV3-008L PE=4 SV=1 | 347 | 44.14 | 2.03 | Hydrophobic | 41% |
Table 1: Representation of various parameters for each protein sequence.
A protein sequence consists of string of amino acids bonded together in a chain. There are 20 amino acids present; the protein sequence is formed by the various combinations of these amino acids. Each amino acid has different nature i.e., they maybe hydrophilic or hydrophobic with respective molecular weight and iso-electric point. Depending on the amount of hydrophobic or hydrophilic amino acids we can determine the nature of the protein sequence. The tool is built on the Java platform. The functionalities of the tool are coded in Java and the interface given to the user is JSP. This can help the user perform particular tasks with ease and there is no necessity for the user to have too much technical knowledge while using this tool.
Results
The tool converts the unstructured dataset into structured dataset. The conversion covers the following functionalities:
Calculations of parameters like molecular weight, point, hydrophobicity, hydrophilicity and neutrality
This product handles the complete Fasta file with all the sequences in it and produces an output in the form of an excel file which becomes a dataset that can be used for further studies. When the user provides a genuine input and selects an appropriate option, the tool computes the parameters such as molecular weight, iso-electric point and classifies the protein sequences into hydrophobic and hydrophilic proteins. The hydrophobic and hydrophilic nature tells us if the protein is soluble in water or not. By this we can conclude if the protein is structural or functional. Table 1 displays the excel file generated.
When the user chooses to generate just one parameter such as molecular weight the tool computes only the molecular weight for the proteins and the output in the excel file is as shown in Table 2.
| Description | Length of the sequence | Molecular weight in KD |
|---|---|---|
| >sp|Q6GZX4|001R_FRG3G Putative transcription factor 001R OS=Frog virus 3 (isolate Goorha) GN=FV3-001R PE=4 SV=1 | 256 | 34.33 |
| >sp|Q6GZX3|002L_FRG3G Uncharacterized protein 002L OS=Frog virus 3 (isolate Goorha) GN=FV3-002L PE=4 SV=1 | 320 | 40.40 |
| >sp|Q197F8|002R_IIV3 Uncharacterized protein 002R OS=Invertebrate iridescent virus 3 GN=IIV3-002R PE=4 SV=1 | 458 | 62.16 |
| >sp|Q197F7|003L_IIV3 Uncharacterized protein 003L OS=Invertebrate iridescent virus 3 GN=IIV3-003L PE=4 SV=1 | 156 | 19.84 |
| >sp|Q6GZX2|003R_FRG3G Uncharacterized protein 3R OS=Frog virus 3 (isolate Goorha) GN=FV3-003R PE=3 SV=1 | 438 | 56.17 |
| >sp|Q6GZX1|004R_FRG3G Uncharacterized protein 004R OS=Frog virus 3 (isolate Goorha) GN=FV3-004R PE=4 SV=1 | 60 | 7.58 |
| >sp|Q197F5|005L_IIV3 Uncharacterized protein 005L OS=Invertebrate iridescent virus 3 GN=IIV3-005L PE=3 SV=1 | 217 | 27.76 |
| >sp|Q6GZX0|005R_FRG3G Uncharacterized protein 005R OS=Frog virus 3 (isolate Goorha) GN=FV3-005R PE=4 SV=1 | 204 | 27.08 |
| >sp|Q91G88|006L_IIV6 Putative KilA-N domain-containing protein 006L OS=Invertebrate iridescent virus 6 GN=IIV6-006L PE=3 SV=1 | 352 | 47.99 |
| >sp|Q6GZW9|006R_FRG3G Uncharacterized protein 006R OS=Frog virus 3 (isolate Goorha) GN=FV3-006R PE=4 SV=1 | 75 | 10.18 |
| >sp|Q6GZW8|007R_FRG3G Uncharacterized protein 007R OS=Frog virus 3 (isolate Goorha) GN=FV3-007R PE=4 SV=1 | 128 | 16.00 |
| >sp|Q197F3|007R_IIV3 Uncharacterized protein 007R OS=Invertebrate iridescent virus 3 GN=IIV3-007R PE=4 SV=1 | 447 | 60.07 |
| >sp|Q197F2|008L_IIV3 Uncharacterized protein 008L OS=Invertebrate iridescent virus 3 GN=IIV3-008L PE=4 SV=1 | 347 | 44.14 |
Table 2: Representation of a specific parameters.
Finding the motif sequences
The user inputs the motif and the application scans for the motif presence that can give further inference where we can identify the kind of function that is performed by the proteins that show a positive result to the motif scan [9]. After the input file name and the motif is entered, the search for motifs starts and the proteins containing the motifs are displayed. (Figure 1) shows the input/output screen shot for this functionality.
Identification of IDP’s
IDPs cover a spectrum of states from fully unstructured to partially structured and include random coils, (pre-) molten globules, and large multi-domain proteins connected by flexible linkers. Proteins are hetero-polymers consisting of covalent linkages between consecutive amino acid monomers [10]. The amino acid sequence of a protein, termed its primary structure, confers chemical properties to the protein through the characteristic properties of the 20 amino acids.
When there is an unusual sequence of amino acids being bonded to form a protein sequence i.e., containing large amount of amino acids with high iso-electric point and a relatively low quantity of hydrophobic amino acids in a protein sequence, the presence of IDP is suspected. From the physical viewpoint, a combination of low hydrophobicity with high net charge represents an obvious prerequisite for intrinsic unfolded high net charge leads to charge-charge repulsion, and low hydrophobicity means less driving force for protein compaction. Here we find those proteins whose hydrophobicity is too low and whose charge is very high. The functionality also provides an option to the user to display the characteristic of the proteins in Graphical and Tabular format.
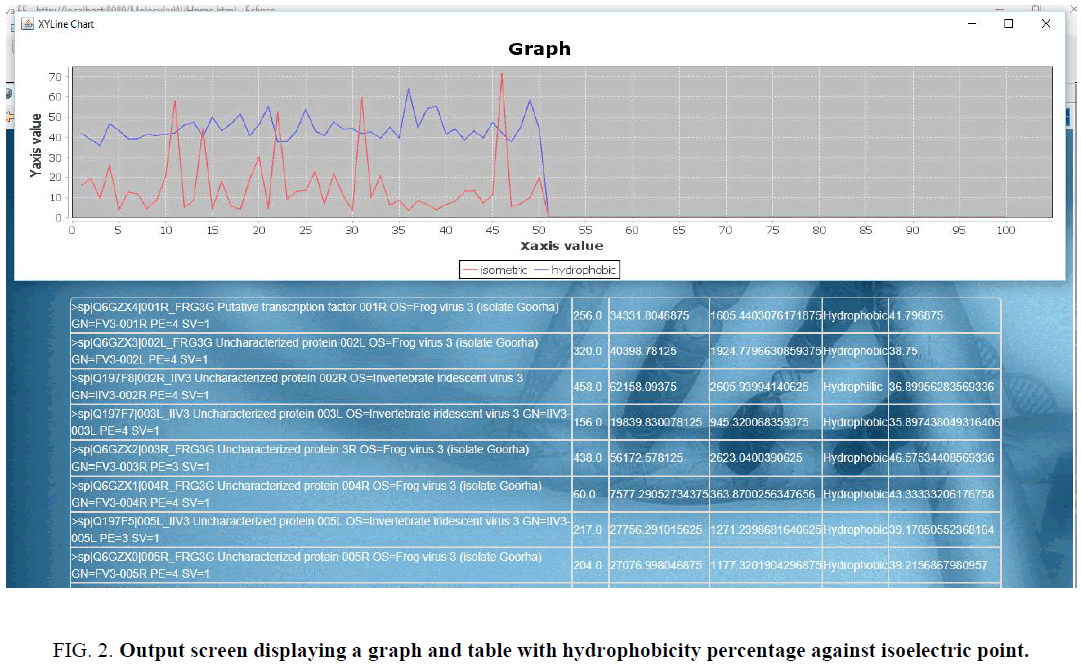
The output generated by the third module – IDP analysis is shown in (Figure 2). User can generate a graph and display the contents of the excel file for analysing the data to find the presence of IDP.
Figure 2: Output screen displaying a graph and table with hydrophobicity percentage against isoelectric point.
Conclusion and Future Scope
In this tool one can efficiently calculate the parameters of proteins such as molecular weight, iso-electric point and classify them into hydrophobic and hydrophilic proteins. The tool accepts a file as its input and provides analyzed output as per the user’s choice. The tool also scans the sequences for presence of a given motif. The user can search for IDPs as well. There is a lot of scope for future enhancements of this tool. The protein mining tool handles a Fasta file of many sequences in it, processes these protein sequences and computes various results. The field of proteomics deals with a huge amount of data which is of the order of giga bytes and above. Thus the tool can be implemented on a Hadoop system to generate faster outputs. The Hadoop Distributed File System (HDFS) is designed to store very large data sets reliably, and to stream those data sets at high bandwidth to user applications. The map-reduce feature of Hadoop processes chunks of data separately and reduces the intermediate outputs to a single output. Hence the calculations can be done for a large number of proteins sequences at a much faster time. This can finally lead to better analytical results.
Acknowledgment
We the authors, are thankful to the Management and Department of Computer Application, M.S. Ramaiah Institute of Technology, Bangalore, Karnataka, India for giving us an opportunity to work on project titled "Protein Mining Tool".
References
- Structural Classification of Proteins. Available from: http://scop.mrc-lmb.cam.ac.uk/scop/
- Marta AB, Robu N. A Study of Sequence Clustering on Protein’s Primary Structure using a Statistical Method. Acta Polytechnica Hungarica. 2006;3(3):17-27.
- National Center for Biotechnology Information. Available from: http://www.ncbi.nlm.nih.gov
- Protein Motifs. Aviable from: http://bioweb.uwlax.edu/genweb/molecular/Seq_Anal/Protein_Motifs/protein_motifs.htm
- Uversky VN. Targeting intrinsically disordered proteins in neurodegenerative and protein dysfunction diseases. Expert Rev Proteomics. 2010;7(4):543-64.
- Saha S, Chaki R. Application of Data Mining in Protein Sequence Classification. Int J Database Manage. 2012;4(5):103-18.
- Yip V, Chen B, Kockara S. Extraction of Protein Sequence Motifs Information by Bi-Clustering Algorithm. BIOCOMP. 2010;185-90.
- Dunker AK, Brown CJ, Obradovic Z. Identification and functions of usefully disordered proteins. Adv Protein Chem. 2002;62:25-49.
- Zhong W, Altun G, Harrison R, et al. Improved K-means clustering algorithm for exploring local protein sequence motifs representing common structural property. IEEE Trans Nanobioscience. 2005;4(3):255-65.
- Radivojac P, Obradovic Z, Smith DK. et al. Protein flexibility and intrinsic disorder. Protein Sci. 2004;13(1):71-80.