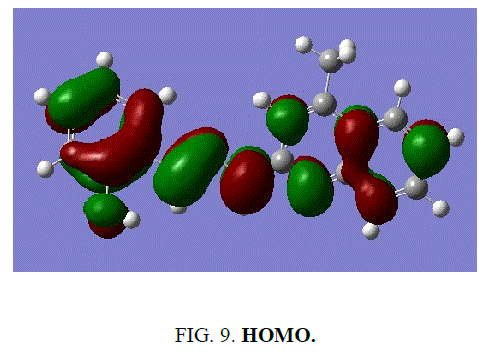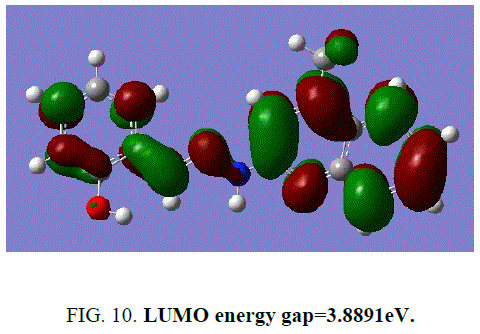Original Article
, Volume: 15( 3)Synthesis, Characterisation, Pharmacological Studies and Quantum Chemical Calculations of Osmium (III) Complex
- *Correspondence:
- Selvi G , Department of Chemistry, PSGR Krishnammal College for Women, Coimbatore, Tamilnadu, India, Tel: 9965152039; E-mail: selvi_gv@rediffmail.com
Received: May 26, 2017; Accepted: July 07, 2017; Published: July 12, 2017
Citation:Sathya Priyadarshini G, Namitha R, Mageswari D, et al. Synthesis, Characterisation, Pharmacological Studies and Quantum Chemical Calculations of Osmium (III) complex. Int J Chem Sci. 2017;15(3):158.
Abstract
New osmium complexes of 4-methyl-2-(salicylidenehydrazino) quinoline H2L1 and 4, 6-dimethyl-2-(salicylidenehydrazino) quinoline H2L2 were synthesised and characterised by spectroscopic method and TGA analysis. Quantum chemical calculations of ligand was performed using the DFT B3LYP method in the 6-31G (d, p) basis. The calculated results are well agreement with the experimental data. The calculated HOMO and LUMO energies show the chemical activity of the molecule. Nonlinear optical behavior of the ligand was analyzed by using first hyperpolarizability. The calculated first hyperpolarisability shows that the ligand is an attractive molecule for future application in nonlinear optics. Based on the theoretical and spectroscopic studies, it has been confirmed that the ligands H2L1 and H2L2 coordinated through phenolate oxygen, ‘N’ of quinoline ring and NH group of hydrazine moiety towards the osmium (III) ion exhibiting octahedral geometry. Antimicrobial result suggests that the complexes having good antimicrobial activity.
Keywords
Ligand; Quinoline; Osmium complex; Drugs
Introduction
Metal complexes with biologically active ligands have attracted more interest from researchers over the last few years. By complexation of these ligands to metal ions, physical and biological activity invitro could be altered significantly [1]. Ligand selection is always an important asset during the design and synthesis of metal complexes. The Chemistry of hydrazinoquinoline derivatives occupy special place because, these ligands developed due to their diverse chelating capability, structural flexibility [2] and pharmacological activities like antibacterial, antifungal, anti-tumoural, antiviral, antimalarial, antituberculosis [3-5]. The use of substituted quinolines are prominent building blocks in both organic and inorganic molecular chemistry with their π-stacking ability and coordination properties. A series of compounds derived from 8-hydroxylquinoline as potential HIV 1 integrase inhibitors were synthesized recently. These compounds show a significant similarity to some novel anti-fungal agents, homoallylamines which possess potent antifungal activity. Another interesting expedition of quinoline derivatives have good nonlinear properties and have been extensively studied due to their great application in the field of organic light emitting diode (OLED). Quinolines have good electron mobility, good thermal and oxidative stability, high photoluminescence efficiency and good film forming properties which is important for they are used in OLED’s [6].
Ruthenium and Osmium-based compounds offer lower toxicity compared to other metals and show different mechanisms of action as well as different spectra of activity compared to Platinum-based drugs [7]. The osmium (III) complexes, particularly the azole chelate ligand, have been widely used in antiproliferative activity [8]. These complexes with pincer ligands have reported as promising alternative to the metal catalyst and are useful due to their photophysical application [9].
The osmium (III) complex has the ability to promote CH activation as well as to accommodate tridenate ligands [10]. The importance of osmium complexes and richness of the quinolines chemistry have now prompted us to explore the reactivity OsCl3.3H2O towards substituted quinolines, which is suited for NNO coordination mode.
Cl3
In our present work, new tridenate Schiff bases have been synthesized by condensing 4-methyl-2-hydrazinoquinoline and 4, 6-dimethyl-2-hydrazinoquinoline with salicylaldehyde. The 1:1 metal complexes were obtained as a result of interaction of ligand and the metal ion Os (III). The IR and NMR spectra of the ligand are reported both theoretically and experimentally. Also, the thermal analysis, Mulliken charge analysis, FMO analysis and first hyperpolarizability is also reported. All the synthesized complexes have been screened for their in vitro biological activities.
Materials and Instruments
Thin layer chromatography (TLC) was performed using glass plates coated with silica gel. Petroleum ether and ethyl acetate was used as eluent. Spots were visualized with iodine. Purification of crude sample was carried out using chromatographic column packed with silica gel. IR spectra were recorded in NICOLET IR 200 FT-IR spectrometer using KBr disc and the absorption frequencies quoted in reciprocal centimeters. UV spectra were recorded in UV-VIS spectrophotometer 3000+, LAB INDIA. 1H NMR spectra were recorded in AMX (300 MHz) spectrometer using tetra methyl silane (TMS) as internal reference.
The chemical shift was expressed in parts per million (ppm). TGA-DSC curves were recorded on Shimadzu 60 H model using 10°C/min. heating rate. The solvents and reagents used for synthesis were of reagents grade and purified by standard methods.
Preparation of Ligand
Preparation of 4-methyl-2-(salicylidenehydrazino) quinoline (3a) (H2L1)
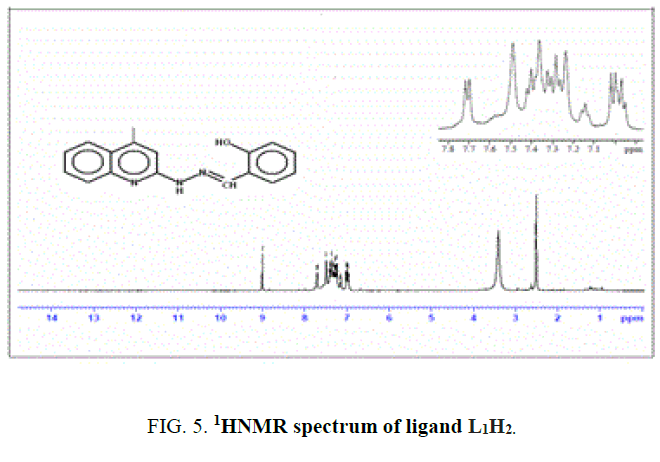
1 mL (6 mmol) of salicyaldehyde (1) was added to 1 g of 4-methyl-2-hydrazinoquinoline (2a) (6 mmol) was dissolved in alcohol. It was refluxed for about 6 h. Completion of the reaction was monitored by TLC. The product was filtered, washed thoroughly with ethanol and dried over vacuum. The compound was recrystallized from ethanol. IR value of ligand υ in cm-1: 3369 (-OH), 3224 (-NH), 1623 (-C=N) and 1251 (-CO).1H NMR value of ligand δ in ppm (300 MHz): 9.01(s, 1H), 6.8 (s, 1H), 7.7-6.9 (m, 8H), 3.4 (s, 1H), 2.4 (s, 3H) Yield-65%, M.P-195°C.
Preparation of 4, 6-dimethyl-2-(salicylidenehydrazino) quinoline (3b) (H2L2)
1 mL (6 mmol) of salicyaldehyde (1) was added to 1.12 g of 4, 6-dimethyl-2-hydrazinoquinoline (2b) (6 mmol) was dissolved in alcohol. It was refluxed for about 6 h. Completion of the reaction was monitored by TLC. The product was filtered, washed thoroughly with ethanol and dried under vacuum. The compound was recrystallized from ethanol. IR value of ligand υ in cm-1: 3289 (-OH), 3050 (-NH), 1608 (-C=N) and 1253 (-CO).1H NMR value of ligand δ in ppm (300 MHz): 9.0 (s, 1H), 6.7 (s, 1H), 7.68-6.98 (m, 8H), 3.4 (s, 1H), 2.4 (s, 3H), 1.2 (s, 3H) Yield – 72%, M.P-205°C.
Preparation of New Osmium (III) Complex [Os (L) Cl (H2O)2]
Preparation of complex A [Os (L1) Cl (H2O)2]
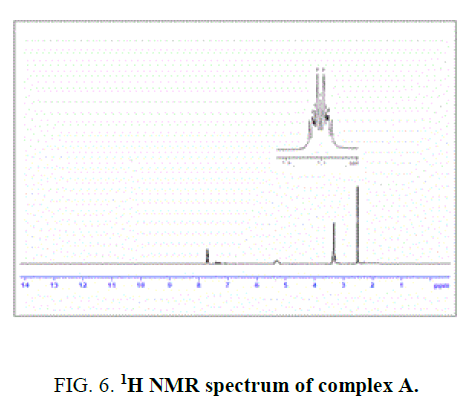
40 mg (0.11 mmol) of osmium trichloridetrihydrate (OsCl3.3H2O) in 20 mL of ethanol was added 30 mg (0.11 mmol) of 4-methyl-2-(salicylidenehydrazino) quinoline (H2L1). The reaction mixture was kept for continuous stirring about 8 h and completion of the reaction was monitored with TLC. After the completion of the reaction, the reaction mixture was allowed to stand overnight. Dark violet crystals separated, washed with ethanol and dried under vacuum. IR value of complexes υ in cm-1: 3361 (-OH, bs), 1628 (-C=N) and 1050 (-CO).1H NMR value of complex δ in ppm (300 MHz): 5.4 (bs, 1H), 7.68-7.75 (m, 8H), 3.4 (s, 1H), 2.4 (s, 3H). Yield-75%, Melting point: >300°C.
Preparation of complex B [Os (L2) Cl (H2O)2]
40 mg (0.11 moles) of osmium trichloridetrihydrate (OsCl3.3H2O) in 20 mL of ethanol was added 30 mg (0.11 mmoles) of 4,6-dimethyl-2-(salicylidenehydrazino) quinoline (H2L2). The reaction mixture was kept for continuous stirring about 8 h and completion of the reaction was monitored with TLC. After the completion of the reaction, the reaction mixture was allowed to stand overnight. Dark blue crystals separated, washed with ethanol and dried under vacuum. IR value of complexes υ in cm-1: 3398 (-OH, bs), 1607(-C=N) and 1201 (-CO).1H NMR value of complex δ in ppm (300 MHz): 5.4 (bs, 1H), 7.68-7.75 (m, 8H), 3.4 (s, 1H), 2.4 (s, 3H), 1.5 (s, 3H). Yield: 75%, Melting point: >300°C.
Antimicrobial activity
The Antibacterial and antifungal activities of Schiff base ligands and complex A and B were studied against the bacteria, Staphylococcus aureus (Sa), Bacillus cereus (Bc), Micrococcus luteus (Ml) and Escherichia coli (Ec) and the fungi, Candida albicans, Aspergillus flavus and Fusarium sp. by Well diffusion method.
Each bacterial strain was suspended in nutrient broth; each fungal strain was suspended in potato dextrose broth and incubated at 37°C. Nutrient Agar (NA) plates were seeded with 8 h broth culture of different bacteria. A 16 h broth culture of Candida albicans, Aspergillus flavus and Fusarium sp was used to seed potato dextrose agar (PDA) plates. In each of these plates, wells were cut out using sterile cork borer. Using sterilized dropping pipettes, different concentrations (25%, 50%, 75% and 100%) of samples were carefully added into the wells and allowed to diffuse at room temperature for 2 h. The plates were then incubated at 37°C for 18 h-24 h. Gentamycin (10 μg) and Clotrimazole (10 μg) were used as positive controls and the respective solvents as negative control. The antimicrobial activity was evaluated by measuring the diameter of inhibition
Quantum chemical calculations
All geometrical optimizations and frequency calculations of the ligand H2L1 were done using the Gaussian 09 program. Density functional theory with the Becke three parameters hybrid functional (DFT-B3LYP) calculations were performed with a 6-31G (d, p) basis set. With the aim to arrive at deeper insight of osmium (III) ion interaction with N, N, O coordination environment, the total energy calculation within the DFT framework were carried out.
Results and Discussion
The present work is focused on the study of coordination behavior of hydrazone Schiff bases (viz.) 4-methyl-2-(salicylidinehydrazino) quinoline and 4,6-dimethyl-2-(salicylidinehydrazino) quinoline with OsCl3. 6H2O by using theoretical and experimental results. All the metal complexes and the ligands were prepared by the molar ratio M: l-1: 1. The complexes are intensively coloured stable solids. The ligand and complexes were soluble in ethanol, chloroform, DMSO and DMF. The structures of the synthesized ligands and the complexes were established from their experimental results (i-e) UV, IR, NMR and TGA analysis and are compared with their theoretical results such as IR and NMR of the ligand, Mulliken charge analysis and FMO study. The formation of the metal complexes can be represented by the following equation and SCHEME 1.
Spectroscopic Studies
Electronic spectra
Solution spectral data of the ligands and Os (III) complexes are collected in TABLE 1.
| Compounds | Molecular weight (g mol-1) | Colour | l in nm | Molar Conductance (mho cm2 mol-1) |
|---|---|---|---|---|
| L1H2 | 277.01 | White | 261, 287, 387 | - |
| L2H2 | 292.03 | Yellow | 246, 290,325 | - |
| [Os (H2O)2 (Cl) L1]. H2O | 536.96 | Violet | 575, 405, 205 | 9 |
| [Os (H2O)2 (Cl) L2]. H2O | 550.99 | blue | 560, 400, 246 | 14 |
TABLE 1. Analytical and physical data of the ligands and the complexes.
The free ligand H2L1 showed three absorption bands at 261 nm, 287 nm and 387 nm. The first band would be assigned to π-π* transition within the aromatic rings. The second band would be due to n-π* transition within ‘C=N’ group. The absorption band at 387 nm is due to charge transfer transition [11,12]. Complex A showed three absorption bands at 575 nm, 405 nm and 205 nm. The band at 575 nm is due to d-d transition.
IR spectra
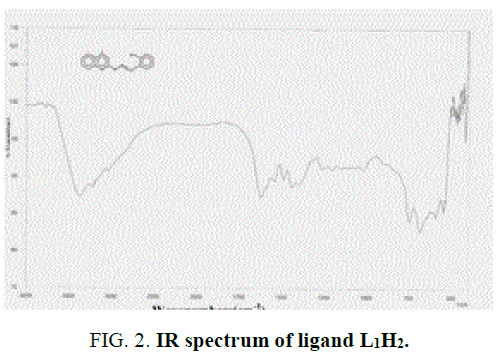
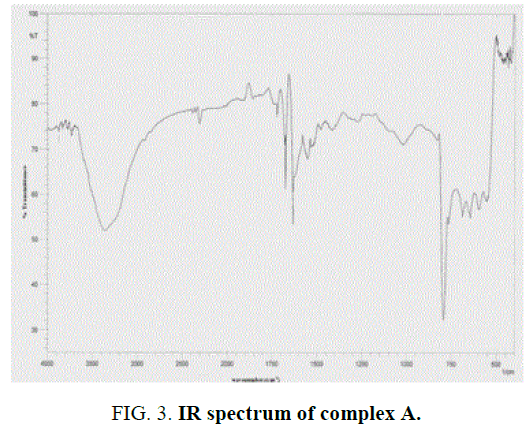
The IR spectra data of the Schiff base ligands (L1H2) (experimentally and theoretically); L2H2 and their complexes are presented in the TABLE 2 and FIG. 2 and FIG. 3 of IR spectra of ligand and complexes. IR spectra of the Schiff base ligand (L1H2) and L2H2 exhibited a strong peak at 1623 cm-1 and 1608 cm-1 was assigned to azomethine group, and for –OH, –NH, C-O stretching vibrations were observed at 3369 cm-1, 3224 cm-1 and 1251 cm-1. IR Spectra of the complex A showed a characteristic peak at 1628 cm-1 and 1050 cm-1 due to stretching vibrations of –C=N and –C=O groups [13].
| Compounds | u (O–H) | u (N-H) | u(C=N) azomethine | u (C–O) |
|---|---|---|---|---|
| L1H2 | 3369 | 3224 | 1623 | 1251 |
| L2H2 | 3289 | 3050 | 1608 | 1253 |
| L2H2 (theoretical) | 3498 | 3213 | 1658 | 1192 |
| [Os (H2O)2 (Cl) L1]. H2O | 3361(bs) | - | 1628 | 1050 |
| [Os (H2O)2 (Cl) L2]. H2O | 3398(bs) | - | 1607 | 1201 |
TABLE 2. Characteristic IR bands of the ligand and the complexes (in cm-1).
The Schiff base ligand has four potential donor sites (i) Quinoline nitrogen (ii) hydrazino nitrogen (iii) azomethine nitrogen (iv) phenolic oxygen. In order to determine the coordination sites, the IR spectra of the complexes were compared with those of the free ligands. Upon comparison, IR value of the metal complex does not show a remarkable shifting frequency of υ (C=N) compared to its respective ligand, this suggests that the nitrogen atom in azomethine group has not participated in the coordination. The broadened band at 3361 cm-1-3400 cm-1 revealed that the presence of coordinated water molecule in the metal complexes.
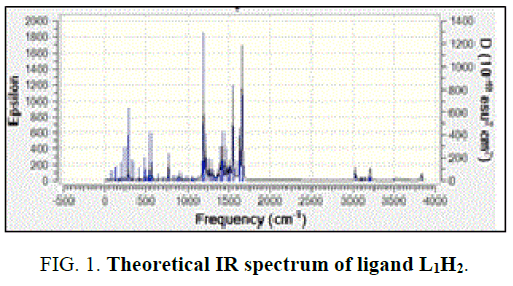
In the free ligand, a strong band at 1200 cm-1 due the C-O (phenolic) shifts to lower frequency in the complexes indicates the coordination of the phenolic oxygen atom to the metal ion. In addition, the change in the intensity of –OH of the IR region shows the disappearance of –OH and of the hydroxyl and appearance of –OH of H2O [14]. Theoretical IR spectra of ligand is shown in FIG. 1. The important assignments (cm-1) are 3498, 3213, 1658 and 1192, these values agree with experimental vibrational frequencies and hence prove the reliability of the applied level of theory [15].
1H NMR spectra
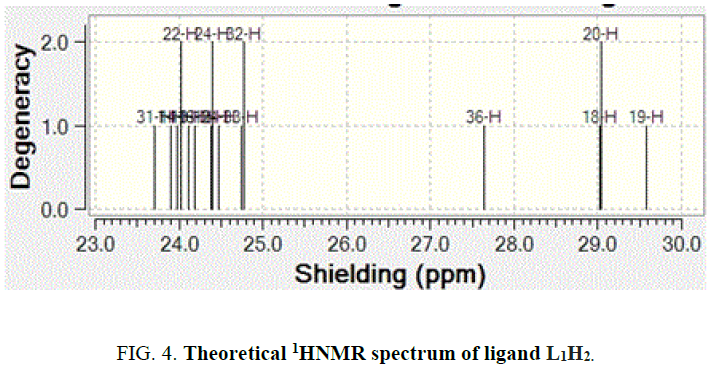
The NMR Spectra of the Schiff bases, H2L1 (both experimentally and theoretically), H2L2 and their Os (III) complexes were recorded in CDCl3 data are given in TABLE 3 and FIG. 4, 5 and 6. 1H NMR Spectra (ppm) of the ligand (L1H2) showed signals δ 9.01 are assignable for –NH protons, δ 7.7-6.9 for aromatic protons, δ 6.7 for –OH protons, δ 3.4 for azomethine protons and δ 2.4 for methyl protons. 1H NMR Spectra (ppm) of the complex A showed signals at δ 7.7-6.9 multiplet for aromatic protons, δ 3.4, singlet for azomethine proton and δ 2.4, singlet for methyl proton [15].
Figure 6: 1H NMR spectrum of complex A.
| Compounds | -NH | -C-OH | Ar–H | -CH=N | -CH3 | -CH3 |
|---|---|---|---|---|---|---|
| L1H2 (3a) | 9.01 | 6.8 | 7.7-6.9 | 3.4 | 2.4 | - |
| L2H2 (3b) | 9.0 | 6.7 | 7.68-6.9 | 3.4 | 2.4 | 1.2 |
| [Os (H2O)2 (Cl) L1]. H2O Complex A |
- | 5.4(bs) | 7.68-7.75 | 3.4 | 2.4 | - |
| [Os (H2O)2 (Cl) L2]. H2O Complex B |
- | 5.4(bs) | 7.68-7.75 | 3.4 | 2.4 | 1.5 |
TABLE 3. 1H-NMR spectral data of the ligand and the Os (III) complexes (in ppm).
The spectra of the complexes are examined in comparison with those of the parent Schiff bases. Upon examination, it was found that –OH and -NH signal that appeared in the spectrum of the H2L1 ligand at 9 ppm and 6.8 ppm completely. Disappeared in the spectrum of its Os (III) complex indicating that the –OH and -NH protons are removed by chelation with Os (III) ion1. The theoretical 1H NMR data of the meaning full region of the ligand well coincides with experimental outcome [16].
Molar conductivity measurements
Molar conductivities of the complexes were measured in DMSO solution within the range of 10-3 concentrations. Both the complexes show molar conductance in the range of 9-15 mho cm2 mol-1 are shown in TABLE 1. These low values indicate the non-electrolytic nature of the complexes [17].
Thermal analysis
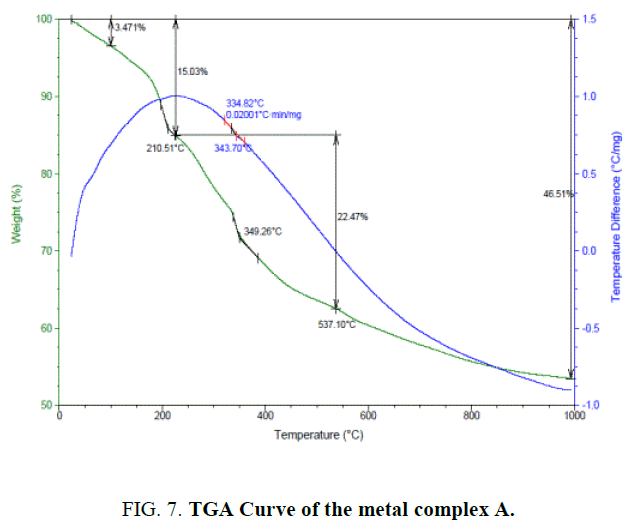
The thermogravimetric studies [18] of the complex A FIG. 7 of the present investigation was carried at the heating rate of 10°C min-1 under nitrogen atmosphere and weight loss as measured from the ambient temperature up to 1000°C. The results obtained for both ligand L1H2 and complex A were tabulated (TABLE 4).
| Compounds | Decomposition Temperature | Weight loss (%) Observed | Cal/ found | Assignments |
|---|---|---|---|---|
| H2 L1 (3a) | 193°C-411.6°C | - | - | Complete decomposition of ligand in one step |
| [Os (H2O)2 L1 (Cl)]. H2O | 110°C 210°C 537°C |
3.47 12.03 22.47 |
3% 12% 22% |
Loss of water molecule in the lattice Loss of coordinated one molecule of water and two chlorine atoms Loss due to the remaining organic moiety |
TABLE 4. TGA data of the ligand H2 L1 and the complex A.
TGA analysis showed that this ligand decomposed into one step from 193.04°C to 411.16°C. This step corresponds to the complete decomposition of H2L1 deduced from mass loss calculation.
The TGA of the metal complex A (FIG. 7) shows three stage decomposition
1. The first decomposition is obtained in the temperature range 110°C. The percentage weight loss (3%) in this range corresponds to the loss of one lattice water molecules.
2. The second stage decomposition is obtained in the temperature range 210.51, (12%) loss indicates the loss of two coordinated water molecules and one chlorine atom.
3. The third stage decomposition is obtained in the temperature range of 400°C-1000°C. The percentage weight loss in the range corresponds to the ligand decomposition and leaving the metal residue.
The residues subsequently fragments of complex A through the several exothermic processes are shown by the DSC curve. (blue color line in FIG. 7).
Mulliken charge analysis
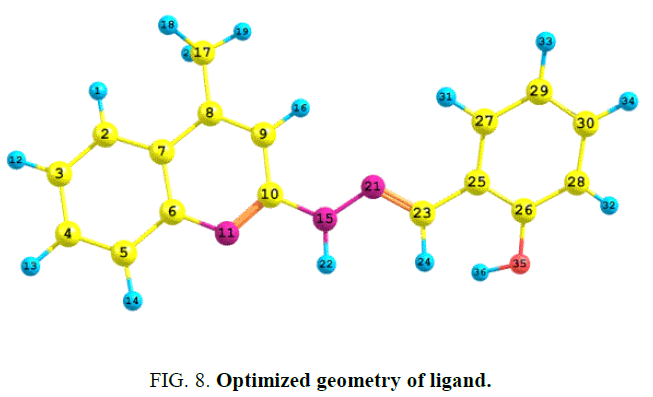
The atomic population and charge distribution in the molecule obtained by the Mulliken analysis and this distribution of the ligand has been calculated by using DFT/B3LYP/6-31G (d, p) level theory [19]. The optimized geometry of the ligand H2L1 was obtained and shown in FIG. 8.
The atomic charges of carbon vary within the range of +0.5277 to -0.1406. From the mulliken charges, it is predicted that the atom N-11(-0.6203) O-35((-0. 5618) and N-15 (-0.4273) acts as a donor site whereas N-21 (-0.2358) is less active towards the coordination.
Frontier molecular orbital analysis
Frontier molecular orbital plays a vital role in the electrical and optical properties as well as chemical reactions. According to FMO of chemical reactivity, transition of electron is due to interactions with other species.
The frontier energy gap of the ligand H2L1 (TABLE 5) is found to be 3.8891eV. The small HOMO-LUMO gap automatically means low excitation energy for the excited states, low stability and highly reactive. The frontier orbitals of the ligand H2 L1 (FIG. 9 and FIG. 10).
| Molecular property | B3LYP/6-31G (d, p) |
|---|---|
| EHOMO | -5.3041 eV |
| ELUMO | -1.4150 eV |
| Energy gap | 3.8891 eV |
TABLE 5. FMO analysis of ligand H2L1.
Figure 10: LUMO energy gap=3.8891eV.
The larger the energy gap, the harder the molecule. The hardness has been associated with the stability of the chemical system. The electron affinity and the global electrophilicity index is also calculated and tabulated in TABLE 6.
| Total Energy after 15 cycles | -896.335722804 A.U. |
|---|---|
| Ionization energy | 5.3041 eV |
| Electron affinity A | 1.4150 eV |
| The global hardness | 1.9445 eV |
| Electronic chemical potential | 3.3565 eV |
| The global electrophilicity index | 2.8969 |
| Dipole moment | 1.1825 |
TABLE 6. Quantum chemical parameters of the ligand H2L1.
Non-linear optical effects
Nonlinear optics deals with the interaction of applied electromagnetic fields in various materials to generate new electromagnetic fields, altered in wavenumber, phase or other physical properties. The calculated first hyperpolarisability of the ligand is 0.427 × 10 ^ -30 e.s.u which is four times that of standard NLO material urea.
Pharmacological activities
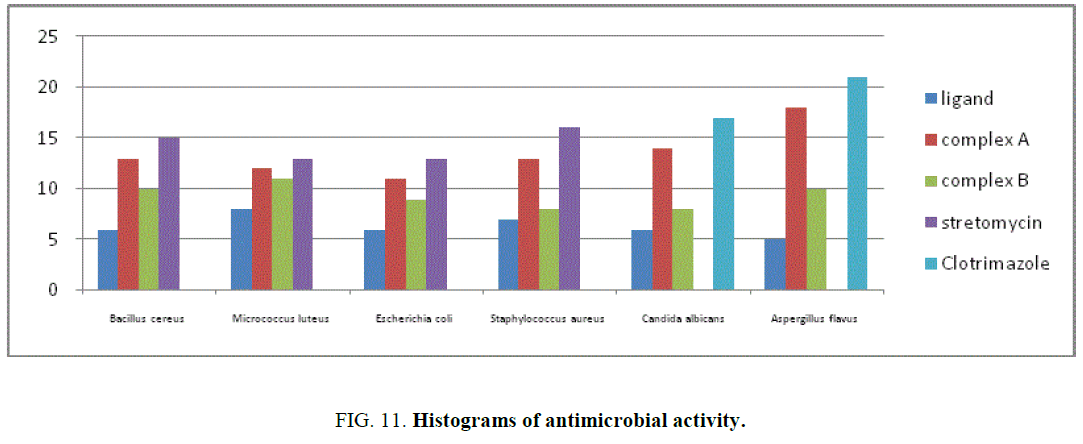
The results of inhibition zone in mm of ligand, complexes A and B for antibacterial and antifungal activity shown in TABLE 7. The histogram of antimicrobial activity was shown in FIG. 11. The antibacterial activity of the ligand, complexes A and B against the bacteria namely Staphylococcus aureus (Sa), Micrococcus luteus (Ml), Bacillus cereus (Bc), Escherichia coli. It was found that the complex was active against Staphylococcus aureus, is almost equal to the standard gentamycin. The complex was less active against Bacillus cereus and Escherichia coli when compared to the standard.
| Compound | Bc | Ml | Ec | Sa | Ca | Af |
|---|---|---|---|---|---|---|
| [Os (H2O)2 (L1) (Cl)]. H2O. Complex A | 12 | 11 | 10 | 13 | 12 | 8 |
| [Os (H2O)2 (L2) (Cl)]. H2O. Complex B | 10 | 11 | 6 | 11 | 11 | 8 |
| Gentamycin | 15 | 13 | 13 | 16 | ||
| Clotrimazole | 17 | 21 |
TABLE 7. Antimicrobial activity of the metal complexes (inhibition zone value in mm).
The enhancement in the activity may be rationalized on the basis that ligands mainly possess C=N bond. The enhanced activity of the complexes over the ligand can be explained on the basis of chelation theory [20]. It is observed that, in a complex, the positive charge of metal is partially shared with the donor atoms present in the ligand and there may be pi electron delocalization over the whole chelation. The increase in the lipophilic character of the metal chelate and favors its permeation through the lipoid layer of the bacterial membranes. The heterocyclic Schiff bases with different fungal groups have greater tendency to interact with nucleoside bases even after complexation with metal ions present in the biosystem can act as promising bactericides because they always tend to interact with enzymatic functional groups, in order to achieve higher coordination number. There are also some other factors which increase the activity namely solubility, conductivity and bond length between the metal and ligand [21].
Conclusion
Osmium complexes having hydrazino quinolines ligands were synthesized from [OsCl3.3H2O] with 4-methyl-2-(salicylidenehydrazino) quinoline (H2L1) and 4, 6-dimethyl-2-(salicylidenehydrazino) quinoline (H2L2). The targeted compound was confirmed by UV, IR,1HNMR spectra, thermal studies and theoretical studies. The theoretical results indicate that mode of co-ordination of the ligand to the metal is N-15, N-21and O-35. The hydrazino quinoline Schiff bases are tridentate and the mode of coordination through NNO (i.e.) phenolate oxygen, nitrogen atom of quinoline and nitrogen atom of hydrazine moiety and suggested geometry as octahedral complexes. From this we conclude that the experimental results of the complex formed is coincide with theoretical result. The synthesized complexes were screened for antibacterial and antifungal activity. The complexes exhibited good antimicrobial activity. The theoretical prediction supports that N-15, N-11 and O-35 of the ligand are involved in the coordination with the metal ion.
References
- Filak LK, Goschl S, Heffler P. Ruthenium and osmium-arene complexes of 2-substituted Indolo [3, 2-c] quinolines: Synthesis, structure, spectroscopic properties, and antiproliferative activity. Organometallics. 2010;32:903-83.
- Rao S, Mishra DD. Oxovanadium binuclear (IV) Schiff base complexes derived from aroyl hydrazones having subnormal magnetic moments. Polyhedron. 1997;16:1825-9.
- West DX, Liberta AE. Thiosemicarbazone complexes of copper(II): Structural and biological studies. Coord Chem Rev. 1993;123:49-71.
- Srivastava A, Chandra A, Singh A. Thiophene-fused quinoline analogues: Facile synthesis of 3-amino-2-cyanothieno [2, 3-b] quinolines from 2-chloro-3-cyanoquinolines. Indian J of Chem. 2005;44:2077.
- Dubey PL, Srinivas Rao S, Aparna V. Synthesis of Some Novel 3-(2-chloro-3-quinolyl)-5-phenyl [1, 3] thiazolo [2, 3-c] [1, 2, 4] triazoles. Heterocyclic communications. 2003;9:281-6.
- Fazal E, Yohannan Panicker C, Tresavarghese H, et al. Spectroscopic investigation (FT-IR, FT-Raman), HOMO–LUMO. NBO analysis and molecular docking study of 4-chlorophenyl quinoline-2-carboxylate. Spectrochimica Acta Part A. 2015;145:260-9.
- Novak MS, Büchel GE, Keppler BK, et al. Biological properties of novel ruthenium- and osmium-nitrosyl complexes with azole heterocycles. J Biol Inorg Chem. 2016;21:347-56.
- Stepaneko IN. Synthesis, structure, spectroscopic properties, and antiproliferative activity in vitro of novel osmium (III) complexes with azole heterocycles. J of Inorg Chem. 2008;47:7338-47.
- Asensio G, Cuenca A, Vlencia M. Osmium (III) complexes with POP pincer ligands: Preparation from commercially available OsCl3·3H2O and their X-ray structures. J of Inorg Chem. 2010;49:8665-7.
- Judith A, Howard K, Sparkes HA, et al. Osmium assisted C–H activation and C double bond; length as m-N cleavage of N-(2′-hydroxyphenyl) benzaldimines. Synthesis, structure and electrochemical properties of some organoosmium complexes. J of Organometallic Chem. 695:2068.
- Mounika K, Anupama B. Synthesis, characterization and biological activity of a Schiff base derived from 3-ethoxy salicylaldehyde and 2-amino benzoic acid and its transition metal complexes. J of Scientific Res. 2010;2:513.
- Abdou S. Synthesis and characterization of chromium (III), molybdenum (III), ruthenium (III) and osmium (III) complexes with N2O2-Schiff bases of 2-hydroxy-1-naphthaldimines. Trans Met Chem. 2004;29:543-9.
- Schmid WF. Highly antiproliferative ruthenium (II) and osmium (II) arene complexes with paullone-derived ligands. Organometallics. 2007;26:6643-52.
- Rajeev TU, Yohannan PC, Hema T, et al. Spectroscopic (FT-IR, FT-Raman) investigations and quantum chemical calculations of 4-hydroxy-2-oxo-1, 2-dihydroquinoline-7-carboxylic acid. Spectrochimica Acta Part A. 2014;121:404-14.
- Kurdekar GS. Synthesis, characterization, antibiogram and DNA binding studies of novel Co (II), Ni (II), Cu (II), and Zn (II) complexes of Schiff base ligands with quinoline core. Med Chem Res. 2011;20:421-9.
- Subramanian N, Sudaraganesan N, Sudha S, et al. Experimental and theoretical investigation of the molecular and electronic structure of anticancer drug camptothecin. Spectrochimica Acta Part A. 2011;8:1058-67.
- Xi PX, Xu ZH. Study on synthesis, structure, and DNA-binding of Ni, Zn complexes with 2-phenylquinoline-4-carboylhydrazide. J of Inorg Biochem. 2009;103:210-8.
- Muhammed GG, Omar MM. Metal complexes of Schiff bases, preparation, characterization and biological activity. Turkish J of Chem. 2006;30:361-82.
- Suliman FO, Al-Nafai I, Al-Busafi SN. Synthesis, characterization and DFT calculation of 4-fluorophenyl substituted tris (8- hydroxyquinoline) aluminum (III) complexes. Spectrochimica Acta Part A. 2014;118:66-72.
- Karekel MR, Biradar V. Synthesis, characterization, antimicrobial, DNA cleavage, and antioxidant studies of some metal complexes derived from Schiff base containing indole and quinoline moieties. J of Bioinorg Chem and App. 2013;2013:1-16.
- Sharma AK, Chandra S. Complexation of nitrogen and sulphur donor Schiff's base ligand to Cr (III) and Ni (II) metal ions: Synthesis, spectroscopic and antipathogenic studies. Spectrochimica Acta Part A. 2011;78:337-42.