Original Article
, Volume: 13( 1)Spectroscopic, physical, thermal and magnetic studies of tricine (L) complexes
- *Correspondence:
- Mohsen M Mostafa , Department of Chemistry, Mansoura University, Mansoura, Dakahlia Governorate 35516, Egypt, Tel: 00-202-01067446662; E-mail: amohsenmostafa@yahoo.com
Received: March 16, 2018; Accepted: March 26, 2018; Published: March 30, 2018
Citation: Hassan EA, Nawar N, Abdel-Galil E, et al. Spectroscopic, Physical, Thermal and Magnetic Studies of Tricine (L) Complexes. Inorg Chem Ind J. 2018;13(1):122
Abstract
New metal complexes derived from the interaction of tricine with some metal salts (Cu2+ and Co2+) were synthesized and characterized by spectral (IR, UV-vis.), magnetic, conductance and thermal analyses (TGA) measurements. The results suggest that L coordinate in bidentate manners via the COO and NH groups. Spectral and magnetic studies suggest an octahedral geometry around the investigated metal ions. Material studio program has been used for calculating HUMO, LUMO and DFT parameters on the atoms to confirm the geometry of complexes.
Keywords
Metal complexes; Spectral and magnetic studies; Metal ions; Solvents; IR spectra; Electronic spectra
Introduction
Tricine (L) exhibits several donor groups [1] and hence it acts as a good chelating agent. Hence, the main goal of our work is to prove that tricine has the flexibility to bind with several metal ions in unidentate, bidentate and/or tridentate manners [2-12]. In continuation of our earlier work [13] and others [14,15] we extend this work to throw the light on the importance of L in different fields with biological studies.
Experimental
Tricine, chemicals and solvents used in the present investigation were of AR quality and used as reported earlier [13]. Also, all instrumentation and (IR, UV-visible), magnetic and physical studies were carried out as reported earlier [13].
Synthesis of complexes in EtOH
A hot EtOH solution of the metal chlorides [CuCl2.2H2O (1.0 mmol, 0.851 g) and CoCl2.6H2O (0.59 g) was added to hot solution of tricine (1.0 mmol, 0.896 g) dissolved in EtOH (25 ml) and few drops of redistilled H2O. The pH of the reaction mixture was adjusted with sodium acetate in case of Cu2+ while in case of Co2+ complexes NaOH was used till the pH of the reaction mixture reaches 7.5. The reaction mixture was refluxed on hot plate for 3 h. The complexes formed were filtered off, washed several times with EtOH and diethyl ether and finally dried in vacuum desiccator over anhydrous CaCl2. The Cu2+ complex is readily soluble in redistilled H2O, DMSO and is partially soluble in EtOH, DMF but the Co2+ complex is partially soluble in H2O, EtOH, DMSO and DMF.
[Co(tric)2Cl2].2H2O
Yield: 90%; brown powder; m. p. ˃ 300 °C. IR (KBr; cm-1): 3415, 3227 [OH (H2O)], OH (EtOH), 2966 (NH), 2880 (OH, acid), 1602 (CO), 521 (M-O). Calcd.: for C12H30CoN2O12Cl2 (%): C.27.5; H, 5.76; Co, 11.24, Cl, 13.52. Found: C, 26.8; H, 5.46; Co, 11.49, Cl, 13.00. Ʌm (DMSO): 8 Ω-1 cm2 mol-1. μeff: 5.79 BM. UV (cm-1): 25252 (LMCT), 18726 (4A2g F → 4T1g P; ν3), 16666 (4A2g → 4T1g; ν2). The values of ν1 (4A2g → 4T2g; ν1), B and β were calculated and found to be 8928 cm-1, 400 cm-1 and 0.41, respectively. The β value indicates that the bond between the L and Co2+ ion is covalent in nature.
[Cu(tric)2]Cl2.3H2O
Yield: 95%; torques powder; m. p. 195 °C. IR (KBr; cm-1): 3322, 3235 [OH (H2O), OH (EtOH), 2895 (OH, acid), 2968 (NH), 1620 (CO), 557 (M-O). Calcd.: for C12H32CuN2O13Cl2 (%): C, 26.35; H, 5.9; Cu, 11.62, Cl,12.96. Found: C, 26.14; H, 5.42; Cu, 11.5, Cl, 13.3. Ʌm (DMSO): 65 Ω-1 cm2 mol-1. μeff: 3.13 BM. UV (cm-1): 31847 cm-1 (LMCT), 12626 cm-1 (2Eg → 2T2g).
Results and Discussion
All the tricine complexes reported earlier in literature [1-15] shows that tricine coordinates to the metal ions in a mononuclear, binuclear and tridentate manner. The results of spectral and magnetic measurements suggest that the ligand coordinates in a bi- and/or tridentate manner and the isolated complexes have an octahedral structure around the metal ions [13].
IR spectra, electronic spectra, magnetic moments and conductance studies
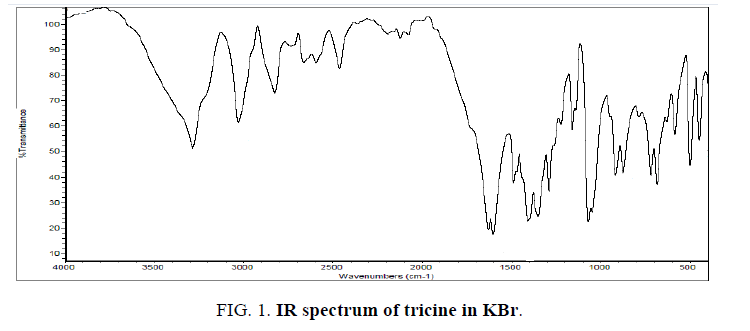
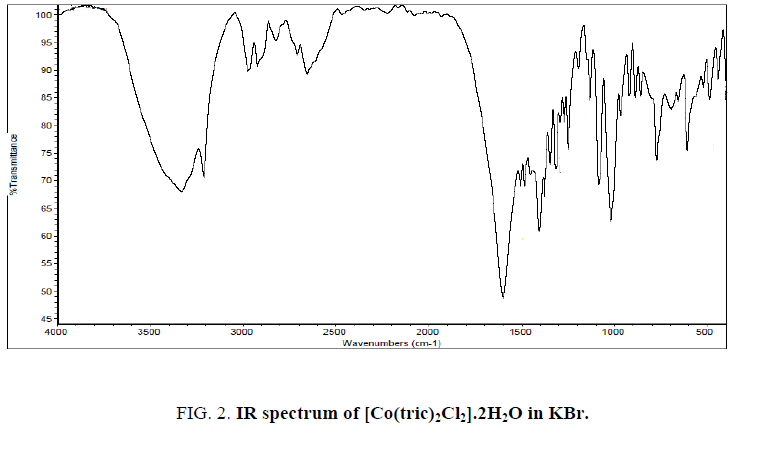
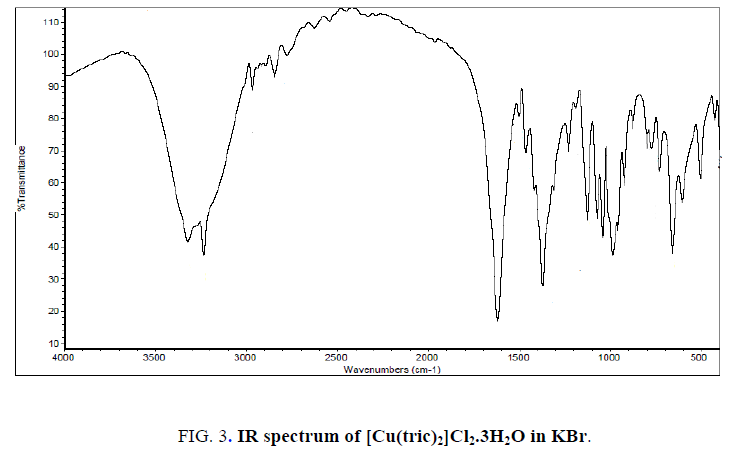
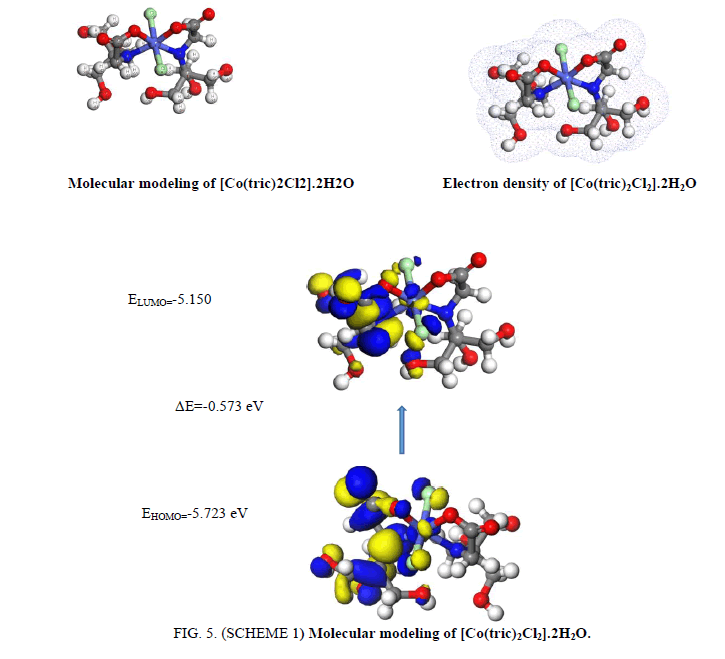
The IR spectrum of tricine (FIG. 1.) was compared with the of the spectra of Co2+, [Co(tric)2Cl2].2H2O and Cu2+ [Cu(tric)2]Cl2.3H2O (FIG. 2 and 3.) complexes. The results indicate that L coordinates to the Co2+ and Cu2+ ions in a bidentate manner through the carboxylate oxygen and amido groups without displacement of a hydrogen atom from the former group [16-18].
The electronic spectra of all complexes were carried out in Nujol mull. The results of electronic spectra as well as the values of magnetic moments indicate that the complexes have octahedral geometry around the metal ions [19-21]. The values of conductance (8-12 ohm-1 cm2 mol-1) confirm that the two complexes are non-electrolyte in nature [22].
Thermal studies
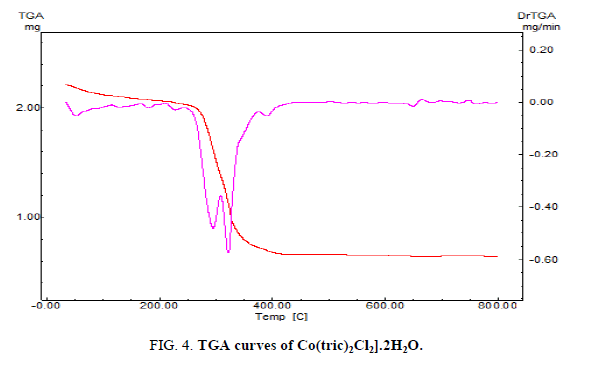
The thermal analyses play a consequential role in studying the properties of the metal complex. To make sure about the proposed formula and the structure of the complex under investigation, the thermal analyses (TGA and DTG) curves was performed under a temperature range from 20 up to 1000 °C. The mass loses were estimated and computed up on the results of the TGA of the calculated mass losses using the results of the microanalyses. The four steps of the decomposition of [Co(tric)2Cl2].2H2O complex is shown in FIG. 4. The temperature of the first step from 25-135 °C corresponds to the loss of two H2O molecules and CH2 (Found: 8.632%, Calcd.: 9.55%). The temperature of the second and third steps from 135-800 °C is referred to the loss of the fragments (C9H22N2O6 +2HCl) (Found: 62.199%, Calcd.: 62.036%). Finally, the residue appraises in the temperature range from 800-1000 °C corresponds to CoNO4C, in which the calculated loss 28.412% which is in matching the found loss 29.1%.
Modeling
The molecular modeling drawing demonstrate the bond lengths, bond angles Tables 1 and 2, chemical reactivity, energy components (Kcal/mol), kinetic energy (Kcal/mol) and binding energy (Kcal/mol) of tricine and its metal complexes are shown in Tables 3 and 4 [23-30]. The DFT theory explains the results [31]. The data of bond angles and lengths illustrate the following comments:
| Bond | Length | Bond | Length |
| O(27)-H(49) | 0.973 | C(20)-C(16) | 1.642 |
| C(24)-H(48) | 1.096 | O(2)-C(25) | 1.305 |
| C(24)-H(47) | 1.094 | C(13)-C(10) | 1.575 |
| C(23)-H(46) | 1.096 | C(6)-O(15) | 1.427 |
| C(23)-H(45) | 1.098 | N(11)-Co(17) | 2.004 |
| O(21)-H(44) | 0.973 | O(26)-C(20) | 1.281 |
| O(19)-H(43) | 0.983 | N(11)-C(7) | 1.442 |
| C(16)-H(42) | 1.091 | C(10)-O(27) | 1.439 |
| C(16)-H(41) | 1.116 | Co(17)-Cl(5) | 2.313 |
| O(15)-H(40) | 0.977 | N(18)-C(12) | 1.433 |
| O(14)-H(39) | 0.973 | C(12)-C(24) | 1.684 |
| C(10)-H(38) | 1.099 | C(24)-O(19) | 1.391 |
| C(10)-H(37) | 1.094 | C(13)-C(6) | 1.597 |
| C(9)-H(36) | 1.099 | N(11)-C(13) | 1.471 |
| C(9)-H(35) | 1.101 | C(25)-C(7) | 1.549 |
| C(7)-H(34) | 1.109 | C(9)-O(21) | 1.431 |
| C(7)-H(33) | 1.095 | C(12)-C(9) | 1.566 |
| C(6)-H(32) | 1.097 | C(20)-O(3) | 1.227 |
| C(6)-H(31) | 1.1 | C(12)-C(4) | 1.546 |
| C(4)-H(30) | 1.099 | C(25)-O(8) | 1.238 |
| C(4)-H(29) | 1.093 | C(13)-C(23) | 1.557 |
| O(1)-H(28) | 0.972 | Co(17)-Cl(22) | 2.304 |
| N(18)-Co(17) | 2.096 | C(16)-N(18) | 1.391 |
| O(26)-Co(17) | 1.975 | C(4)-O(14) | 1.444 |
| O(2)-Co(17) | 1.941 | C(23)-O(1) | 1.426 |
| Bond | Angle | Bond | Angle |
| H(49)-O(27)-C(10) | 107.405 | C(10)-C(13)-N(11) | 107.333 |
| Co(17)-O(26)-C(20) | 114.468 | C(10)-C(13)-C(23) | 110.705 |
| O(2)-C(25)-C(7) | 114.852 | C(6)-C(13)-N(11) | 112.178 |
| O(2)-C(25)-O(8) | 124.412 | C(6)-C(13)-C(23) | 103.403 |
| C(7)-C(25)-O(8) | 120.72 | N(11)-C(13)-C(23) | 114.405 |
| H(48)-C(24)-H(47) | 112.377 | N(18)-C(12)-C(24) | 106.009 |
| H(48)-C(24)-C(12) | 103.962 | N(18)-C(12)-C(9) | 115.038 |
| H(48)-C(24)-O(19) | 114.014 | N(18)-C(12)-C(4) | 112.236 |
| H(47)-C(24)-C(12) | 106.823 | C(24)-C(12)-C(9) | 104.343 |
| H(47)-C(24)-O(19) | 107.81 | C(24)-C(12)-C(4) | 107.85 |
| C(12)-C(24)-O(19) | 111.648 | C(9)-C(12)-C(4) | 110.693 |
| H(46)-C(23)-H(45) | 108.91 | Co(17)-N(11)-C(7) | 105.713 |
| H(46)-C(23)-C(13) | 107.941 | Co(17)-N(11)-C(13) | 137.971 |
| H(46)-C(23)-O(1) | 106.774 | C(7)-N(11)-C(13) | 116.09 |
| H(45)-C(23)-C(13) | 106.233 | H(38)-C(10)-H(37) | 108.87 |
| H(45)-C(23)-O(1) | 110.318 | H(38)-C(10)-C(13) | 108.941 |
| C(13)-C(23)-O(1) | 116.48 | H(38)-C(10)-O(27) | 109.943 |
| H(44)-O(21)-C(9) | 106.118 | H(37)-C(10)-C(13) | 109.487 |
| C(16)-C(20)-O(26) | 112.841 | H(37)-C(10)-O(27) | 110.726 |
| C(16)-C(20)-O(3) | 116.544 | C(13)-C(10)-O(27) | 108.847 |
| O(26)-C(20)-O(3) | 130.612 | H(36)-C(9)-H(35) | 107.241 |
| H(43)-O(19)-C(24) | 106.149 | H(36)-C(9)-O(21) | 111.301 |
| Co(17)-N(18)-C(12) | 137.244 | H(36)-C(9)-C(12) | 111.395 |
| Co(17)-N(18)-C(16) | 102.656 | H(35)-C(9)-O(21) | 110.969 |
| C(12)-N(18)-C(16) | 116.821 | H(35)-C(9)-C(12) | 105.469 |
| N(18)-Co(17)-O(26) | 80.109 | O(21)-C(9)-C(12) | 110.278 |
| N(18)-Co(17)-O(2) | 166.821 | H(34)-C(7)-H(33) | 107.953 |
| N(18)-Co(17)-N(11) | 110.215 | H(34)-C(7)-N(11) | 109.124 |
| N(18)-Co(17)-Cl(5) | 95.382 | H(34)-C(7)-C(25) | 103.551 |
| N(18)-Co(17)-Cl(22) | 90.897 | H(33)-C(7)-N(11) | 114.731 |
| O(26)-Co(17)-O(2) | 86.741 | H(33)-C(7)-C(25) | 112.312 |
| O(26)-Co(17)-N(11) | 169.61 | N(11)-C(7)-C(25) | 108.542 |
| O(26)-Co(17)-Cl(5) | 88.278 | H(32)-C(6)-H(31) | 108.079 |
| O(26)-Co(17)-Cl(22) | 88.107 | H(32)-C(6)-O(15) | 106.974 |
| O(2)-Co(17)-N(11) | 82.951 | H(32)-C(6)-C(13) | 108.727 |
| O(2)-Co(17)-Cl(5) | 85.039 | H(31)-C(6)-O(15) | 109.802 |
| O(2)-Co(17)-Cl(22) | 87.761 | H(31)-C(6)-C(13) | 105.967 |
| N(11)-Co(17)-Cl(5) | 89.532 | O(15)-C(6)-C(13) | 117.015 |
| N(11)-Co(17)-Cl(22) | 92.776 | H(30)-C(4)-H(29) | 110.16 |
| Cl(5)-Co(17)-Cl(22) | 172.115 | H(30)-C(4)-C(12) | 107.422 |
| H(42)-C(16)-H(41) | 113.087 | H(30)-C(4)-O(14) | 109.196 |
| H(42)-C(16)-C(20) | 110.527 | H(29)-C(4)-C(12) | 109.798 |
| H(42)-C(16)-N(18) | 118.296 | H(29)-C(4)-O(14) | 110.485 |
| H(41)-C(16)-N(18) | 109.176 | Co(17)-O(2)-C(25) | 113.811 |
Table 1: Bond lengths (Å) and bond angles of [Co(tric)2Cl2].2H2O using DFT-method from DMOL3calculations.
| Bond | Length(Å) | Bond | Length(Å) | ||
| O(24)-H(45) | 0.97 | C(4)-C(7) | 1.589 | ||
| O(23)-H(44) | 0.972 | C(4)-C(11) | 1.549 | ||
| C(21)-H(43) | 1.105 | C(11)-O(6) | 1.406 | ||
| C(21)-H(42) | 1.108 | N(19)-C(4) | 1.461 | ||
| O(18)-H(41) | 1 | C(16)-O(2) | 1.255 | ||
| C(17)-H(40) | 1.11 | O(3)-Cu(22) | 2.237 | ||
| C(17)-H(39) | 1.102 | C(20)-C(9) | 1.542 | ||
| C(15)-H(38) | 1.103 | N(19)-Cu(22) | 1.918 | ||
| C(15)-H(37) | 1.105 | C(20)-C(17) | 1.614 | ||
| C(12)-H(36) | 1.097 | C(21)-N(14) | 1.435 | ||
| C(12)-H(35) | 1.101 | C(1)-O(18) | 1.346 | ||
| C(11)-H(34) | 1.106 | C(16)-C(21) | 1.544 | ||
| C(11)-H(33) | 1.102 | C(13)-O(25) | 1.236 | ||
| C(9)-H(32) | 1.099 | O(6)-Cu(22) | 1.913 | ||
| C(9)-H(31) | 1.102 | C(20)-C(1) | 1.792 | ||
| C(7)-H(30) | 1.096 | C(13)-C(15) | 1.544 | ||
| C(7)-H(29) | 1.1 | O(10)-Cu(22) | 2.015 | ||
| O(5)-H(28) | 0.971 | N(14)-C(20) | 1.352 | ||
| C(1)-H(27) | 1.097 | O(10)-C(13) | 1.321 | ||
| C(1)-H(26) | 1.093 | O(8)-Cu(22) | 2.154 | ||
| N(14)-Cu(22) | 2 | C(12)-O(24) | 1.435 | ||
| C(15)-N(19) | 1.452 | C(17)-O(8) | 1.349 | ||
| C(4)-C(12) | 1.571 | O(3)-C(16) | 1.281 | ||
| C(9)-O(5) | 1.43 | C(7)-O(23) | 1.421 | ||
| Angle | Degree (°) | Angle | Degree (°) | Angle | Degree (°) |
| H(45)-O(24)-C(12) | 107.274 | C(15)-N(19)-Cu(22) | 110.1 | H(34)-C(11)-H(33) | 108.38 |
| H(44)-O(23)-C(7) | 107.774 | C(4)-N(19)-Cu(22) | 111.597 | H(34)-C(11)-C(4) | 111.689 |
| N(14)-Cu(22)-O(3) | 78.439 | H(41)-O(18)-C(1) | 111.206 | H(34)-C(11)-O(6) | 107.303 |
| N(14)-Cu(22)-N(19) | 175.925 | H(40)-C(17)-H(39) | 108.797 | H(33)-C(11)-C(4) | 108.087 |
| N(14)-Cu(22)-O(6) | 93.485 | H(40)-C(17)-C(20) | 101.092 | H(33)-C(11)-O(6) | 111.947 |
| N(14)-Cu(22)-O(10) | 96.349 | H(40)-C(17)-O(8) | 113.132 | C(4)-C(11)-O(6) | 109.468 |
| N(14)-Cu(22)-O(8) | 80.029 | H(39)-C(17)-C(20) | 109.548 | Cu(22)-O(10)-C(13) | 111.882 |
| O(3)-Cu(22)-N(19) | 97.519 | H(39)-C(17)-O(8) | 111.415 | H(32)-C(9)-H(31) | 107.672 |
| O(3)-Cu(22)-O(6) | 92.302 | C(20)-C(17)-O(8) | 112.35 | H(32)-C(9)-O(5) | 111.276 |
| O(3)-Cu(22)-O(10) | 93.608 | O(2)-C(16)-C(21) | 118.132 | H(32)-C(9)-C(20) | 110.103 |
| O(3)-Cu(22)-O(8) | 158.415 | O(2)-C(16)-O(3) | 124.633 | H(31)-C(9)-O(5) | 111.469 |
| N(19)-Cu(22)-O(6) | 86.114 | C(21)-C(16)-O(3) | 117.235 | H(31)-C(9)-C(20) | 108.167 |
| N(19)-Cu(22)-O(10) | 84.361 | H(38)-C(15)-H(37) | 105.805 | O(5)-C(9)-C(20) | 108.113 |
| N(19)-Cu(22)-O(8) | 104.025 | H(38)-C(15)-N(19) | 109.831 | Cu(22)-O(8)-C(17) | 108.833 |
| O(6)-Cu(22)-O(10) | 169.371 | H(38)-C(15)-C(13) | 108.583 | H(30)-C(7)-H(29) | 108.813 |
| O(6)-Cu(22)-O(8) | 90.486 | H(37)-C(15)-N(19) | 114.864 | H(30)-C(7)-C(4) | 106.806 |
| O(10)-Cu(22)-O(8) | 87.262 | H(37)-C(15)-C(13) | 107.054 | H(30)-C(7)-O(23) | 106.545 |
| H(43)-C(21)-H(42) | 105.57 | N(19)-C(15)-C(13) | 110.439 | H(29)-C(7)-C(4) | 108.822 |
| H(43)-C(21)-N(14) | 111.465 | Cu(22)-N(14)-C(21) | 117.306 | H(29)-C(7)-O(23) | 111.922 |
| H(43)-C(21)-C(16) | 108.736 | Cu(22)-N(14)-C(20) | 118.651 | C(4)-C(7)-O(23) | 113.689 |
| H(42)-C(21)-N(14) | 110.122 | C(21)-N(14)-C(20) | 123.932 | C(11)-O(6)-Cu(22) | 107.119 |
| H(42)-C(21)-C(16) | 108.257 | O(25)-C(13)-C(15) | 122.404 | H(28)-O(5)-C(9) | 106.737 |
| N(14)-C(21)-C(16) | 112.413 | O(25)-C(13)-O(10) | 122.064 | C(12)-C(4)-C(7) | 107.444 |
| C(9)-C(20)-C(17) | 117.697 | C(15)-C(13)-O(10) | 115.263 | C(12)-C(4)-C(11) | 111.465 |
| C(9)-C(20)-C(1) | 104.649 | H(36)-C(12)-H(35) | 109.266 | C(12)-C(4)-N(19) | 113.885 |
| C(9)-C(20)-N(14) | 117.588 | H(36)-C(12)-C(4) | 107.815 | C(7)-C(4)-C(11) | 112.783 |
| C(17)-C(20)-C(1) | 100.397 | H(36)-C(12)-O(24) | 111.273 | C(7)-C(4)-N(19) | 109.955 |
| C(17)-C(20)-N(14) | 107.407 | H(35)-C(12)-C(4) | 108.883 | C(11)-C(4)-N(19) | 101.363 |
| C(1)-C(20)-N(14) | 107.126 | H(35)-C(12)-O(24) | 109.003 | Cu(22)-O(3)-C(16) | 113.028 |
| C(15)-N(19)-C(4) | 121.272 | C(4)-C(12)-O(24) | 110.558 | H(27)-C(1)-H(26) | 114.644 |
Table 2: Bond lengths (Å) and bond angles of [Cu(tric)2]Cl2.3H2O using DFT-method from DMOL3calculations.
| Compound | EH eV | EL eV | (EHEL) eV | ᵡ eV | µ eV | Ƞ eV | Ѕ eV-1 | ω eV | σ eV |
|---|---|---|---|---|---|---|---|---|---|
| L1; C6H13NO5 | -5.18 | -0.678 | -4.502 | 2.929 | -2.929 | 2.251 | 1.1255 | 1.9056 | 0.4442 |
| [Co(L1)2Cl2].2H2O | -5.723 | -5.15 | -0.573 | 5.437 | -5.4365 | 0.287 | 0.14325 | 51.5803 | 3.49 |
| [Cu(L1)2]Cl2.3H2O | -5.278 | -4.958 | -0.32 | 5.118 | -5.118 | 0.16 | 0.08 | 81.856 | 6.25 |
Table 3: Calculated E HOMO, ELUMO, energy band gap (EH-EL), chemical potential (μ), electronegativity (ᵡ), global hardness (Ƞ), global softness (Ѕ) and global electrophilicity index (ω) for tricine and its complexes.
| No. | Compound | HOMO (eV) | LUMO (eV) | Binding energy (Kcal/mol) | Total energy (Kcal/mol) |
|---|---|---|---|---|---|
| 1 | L1; C6H13NO5 | -5.18 | -0.678 | -2483726 | -4.189 × 105 |
| 2 | [Co(L1)2Cl2].2H2O | -5.723 | -5.15 | -4863.411 | -1.518 × 106 |
| 3 | [Cu(L1)2]Cl2.3H2O | -5.278 | -4.958 | -4510.29 | -9.751 × 105 |
Table 4: Some of energetic properties of tricine and its complexes calculated by DMOL3 using DFT-method.
1. The [Co(tric)2Cl2].2½H2O complex (FIG. 5. SCHEME 1) where the cobalt atom arranges in octahedral geometry. The ligand of two tricine acts as a bi-dentate manner coordinating via N(18), O(26), O(2) and N(11), additionally two chloride ions Cl(5) and Cl(22) complete the octahedral structure. There is a huge change in N(18)-C(26), N(11)-C(13), O(26)-C(20) and O(2)-C(25). The bond angles of tricine are adjusted upon coordination. The bond angles of tricine are lessened or expanded on complex arrangement as a result of bonding such as C(12)-N(18)-C(16), C(10)_C(13)-N(11), C(26)-C(20)-O(3) and C(6)-C(13)-N(11).
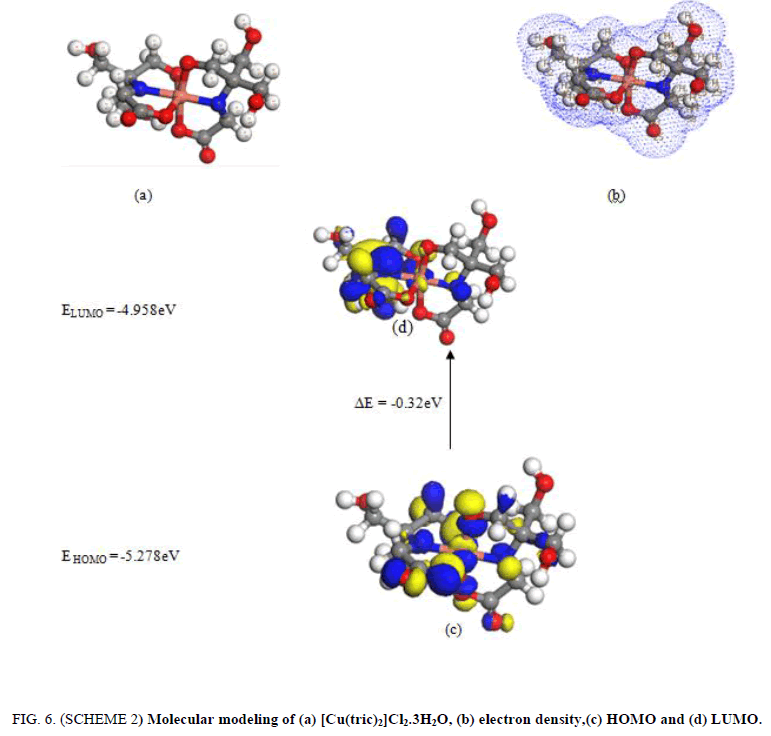
2. The [Cu(tric)2].Cl2.3H2O (FIG. 6. SCHEME 2) has an octahedral structure. The two ligand of tricine act as a tridentate chelating via O(6), O(8), N(14), N(19),O(4) and O(10). The bonds of all active groups joining in coordination are longer than that of now exist ligand such as NH. There is a huge variety in N(19)-C(4), N(14)-C(21),O(8)-C(17), O(6)-C(11), O(3)-C(16) and O(10)-C(13) bond lengths. The bond angles are diminished or expanded on complex arrangement as an outcome of bonding such as C(21)-N(14)-C(13), C(4)-C(11)-O(6).
Figure 6: (SCHEME 2) Molecular modeling of (a) [Cu(tric)2]Cl2.3H2O, (b) electron density,(c) HOMO and (d) LUMO.
Chemical reactivity
The assignment of energies of HOMO (π-donor) and LUMO (π-acceptor) are important parameters in quantum compound counts. The HOMO is the orbital that generally goes about as electron giver and the LUMO is the orbital that fundamentally goes about as electron acceptor. These molecular orbitals are also called frontier molecular orbitals (FMOs). The all negative value of EHOMO, ELUMO and their neighboring orbitals show that the prepared molecules are steady. The energy gap (EHOMO-ELUMO) is an important stability index serves to portray the chemical reactivity and kinetic stability of the molecule [31]. The gap (EHOMO-ELUMO) is connected to build up a hypothetical pattern for illustrating the structure in many molecular systems. The small gap in molecule means the molecule is more polarized and the molecule is known as soft molecule. The responsive of soft molecule is more than hard one as they easily offer electrons to an acceptor. The small energy gap in tricine shows that charge transfer easily happens in it. The ability of the molecule to give electron is weaker if the HOMO energy value is low. On opposite, the ability of the molecule is good if the HOMO energy value is high [31]. All the data are shown in TABLES 3 and 4.
Conclusion
The Cu2+ and Co2+ complexes coordinate in bidentate fashion. The geometry of the isolated complexes was determined by chemical, physical, spectral, conductance and magnetic measurements. The results of spectral and magnetic studies suggest octahedral geometry around the metal ions. The molecular modeling drawing demonstrate the bond lengths, bond angles, chemical reactivity, energy components (Kcal/mol), kinetic energy (Kcal/mol) and binding energy (Kcal/mol) of tricine and its metal complexes.
References
- Crans DC, Ehde PM, Shin PK, et al. Structural and kinetic characterization of simple complexes as models for vanadate-protein interactions. J Am Chem Soc. 1991;113:3728-36.
- Chakraborty D, Bhattacharya PK. Intramolecular interligand interactions in Cu (II) ternary complexes involving dipeptides and amino acids. J Inorg Biochem. 1990;39:1-8.
- Manjula V, Chakraborty D, Bhattacharya P. Ternary complexes of Cu(II) involving histidine and another amino acid or dipeptide. Indian J Chem. 1990;29A:577-80.
- Kapoor RC, Jailwal JK, Kishan JA. Complex Formation of N-[Tris (hydroxymethyl) methyl] glycine with Lead and Cadmium. J Inorg Nucl Chem. 1978;40:155-8.
- Tripathi RM, Ghose R, Ghose AK. Heteroligand complexes of some transition-metals containing 2, 2'-bipyridyl and tricine as ligands in aqueous-solution. Indian J Chem. 1985;24A:565-7.
- Vaidyan AV, Bhattacharya PK. Interligand interaction in ternary complexes of Zn (II) and Cd (II) with dipeptides and aminoacids. Can J Chem. 1994;72:1107-10.
- de Castro B, Pereira J, Gameiro P, et al. Multinuclear NMR and potentiometric studies on the interaction of zinc and cadmium with cytidine and glycylglycine. The effect of the anion. J Inorg Biochem. 1992;45:53-64.
- Pandeya KB, Patel RN. Ternary complexes of copper(II) with glycylglycine, glycylglyclglycine and some imidazoles. Indian J Chem. 1990;29A:602-4.
- Kishan JAI, Kapoor R. Complex Formation of N-[Tris (hydroxy-methyl)methyl]glycine with Copper(II) and Zinc(II). Indian J Chem. 1984;23A:355-6.
- Farkas E, Kiss T. Effects of side-chain donor groups on deprotonation of peptide amide in copper (II) complexes at high pH. Polyhedron. 1989;8:2463-7.
- Hosny WM. Complexes of Vitamin B6. Ternary complexes of Cu(II) with pyridoxamine and amino acids, DNA units or peptides. Egypt J Chem. 1999;42:151-73.
- Prenesti E, Daniele PG, Prencipe M, et al. Spectrum-structure correlation for visible absorption spectra of copper (II) complexes in aqueous solution. Polyhedron. 1999;18:3233-41.
- Al-Radadi NS, Al-Ashqar SMA, Mostafa MM. Synthesis and characterization of some new binary and ternary CuII complexes. Synth React Inorg Met-Org and Nano Chem. 2011;41: 203-10.
- Boraei AA, Ahmed IT. Synth React Inorg Met-Org and Nano Chem. 2002;32: 981-1000.
- El-Roudi OM, Abd Alla EM, Ibrahim SA. Potentiometric studies on the binary complexes of N-[Tris (hydroxymethyl) methyl] glycine with Th4+, Ce3+, La3+, and UO22+ and medium effects on a Th- tricine binary complex. J Chem Eng Data. 1997;42:609-13.
- Nakamoto K. Infrared and Raman spectra of inorganic and coordination compounds Part A. Theory and Applications in Inorganic Chemistry. 1997:88-97.
- Ketcham KA, Garcia I, Swearingen JK, et al. Spectral studies and X-ray crystal structures of three nickel (II) complexes of 2-pyridineformamide 3-piperidylthiosemicarbazone. Polyhedron. 2002;21:859-65.
- Bates RG, Roy RN, Robinson RA. Solute-solvent effects in the acidic dissociation of the ampholyte N-tris (hydroxymethyl) methylglycine (“tricine”) in 50 mass% methanol-water solvent. J Sol Chem. 1974;3:905-16.
- Lever ABP. Inorganic electronic spectroscopy. Amsterdam, The Netherlands. 1968:420.
- Tanabe Y, Sugano S. On the absorption spectra of complex ions I and II. J Phys Soc Jpn. 1954;9:753-66.
- Earnshaw A. Introduction to Magnetochemistry. Acadmic Press. London. 1968.
- Geary WJ. The use of conductivity measurements in organic solvents for the characterisation of coordination compounds. Coord Chem Revs. 1971;7:81-122.
- Kivelson D, Neiman R. ESR studies on the bonding in copper complexes. J Chem Phys. 1961;35:149-55.
- Singh B, Yadava BP, Aggarwal RC. Indian J Chem. 1984;23A:441-52.
- Delley B. An all-electron numerical method for solving the local density functional for polyatomic molecules. J Chem Phys. 1990;92:508-17.
- Delley B. A scattering theoretic approach to scalar relativistic corrections on bonding. Int J Quantum Chem. 1998;69:423-33.
- Delley B. From molecules to solids with the DMol 3 approach. J Chem Phys. 2000;113:7756-64.
- Wu X, Ray AK. Density-functional study of water adsorption on the PuO 2 (110) surface. Phys Rev. 2002;65:085403.
- Kessi A, Delley B. Density functional crystal vs. cluster models as applied to zeolites. Int J Quant Chem. 1998;68:135-44.
- Materials Studio v 5.0. Accelrys software Inc. 2009.
- Linert W, Taha A. Co-ordination of solvent molecules to square-planar mixed-ligand nickel(II) complexes: A thermodynamic and quantum-mechanical study. J Chem Soc Dalton Trans. 1994:1091-5.