Editorial
, Volume: 16( 2)Synthesis, Characterization of Nanosized Copper(II) Complex and its Antimicrobial Activities
- *Correspondence:
- Mbonu Idongesit Justina , Department of Chemistry, College of Science, Federal University of Petroleum Resources, Effurun, Delta State, Nigeria, E-Mail: idongesitmbonu@yahoo.com
Received: August 7, 2018; Accepted: August 9, 2018; Published: August 16, 2018
Citation: Justina MI, Olukemi BAO, Sunday O. Study on “Synthesis, Characterization of Nanosized Copper(II) Complex and its Antimicrobial Activities”. ChemXpress. 2018;11(2):136.
Abstract
The synthesis and characterization of nano-sized Copper(II) complex with succinic acid and DL alanine is reported using elemental analysis, magnetic susceptibility, molar conductance, FT-IR, UV-Vis spectral and powder X-ray diffraction studies. The molar conductance value of 2.2×103 ÃŽÂÂ-1 cm2mol-1 measurement of the complex in DMF solution corresponds to non-electrolytic nature. The complex was found to be high-spin type and exhibited octahedral geometry. The two fungi (Candida albicans and Aspergillus niger) and three bacteria (Klebsiella spp, Shigellaspp, Salmonella specie) profile for synthesized copper(II) complex, ligands and standard antibiotics were investigated. The metal complex showed remarkable antimicrobial activity than some of the recommended antibacterial and antifungal standards. The Copper(II) complex displayed enhanced antifungal activity when compared with the standard drug, fluconazol and free ligands. This present findings may open a new search for this complex to be used in treatment of fungal diseases.
Keywords
Copper (II); Characterization; Antimicrobial activities.
Introduction
The resistances of bacteria and fungi have reduced the efficiency of antimicrobial therapies with the current antibiotics; this challenge has prompted the scientific community to sort for more efficient drugs for the treatment of infectious diseases [1-3]. Research has shown that metals coordinated to biologically active molecules have enhanced biological activity and are used in combating pathogenic microorganisms by encapsulating the microbes and thus restrict their further growth [1-7]. It has been observed that dugs administered in the form of metallic complexes exhibits modified pharmacological and toxicological properties; example platinum and Cu2+ complexes drugs have efficient bioactivity against diseases such as cancer, gastric ulcers and rheumatoid arthritis and cancer [4,6]. In this regard, mixed ligand complexes are of great importance in biological and industrial fields. Substantial researches in chemistry have been devoted to the syntheses and characterization of biologically active compounds that would be used to curb organisms causing illnesses and death to both plants and animals [1-5]. Succinic acid and DL-alanine, in addition to their chemical uses, possess medicinal properties like, antiHIV, antitumor, antiseptic, antioxidant, antifungal and antibacterial activities [4,6]. As part of the ongoing research on mixed ligand complexes, we herein report the synthesis, characterization and biological activity of Copper(II) ions with succinic acid and DL-alanine to investigate if this compound can inhibit the growth of harmful microbes. For the characterization of the compound the following spectroscopic and analytical techniques were used: elemental analysis, FT-IR spectroscopy, UV-Visible, magnetic moment, scanning electron microscopy, powder X-ray diffraction study and magnetic susceptibility measurement. The biological activity of the ligands and their metal complex is evaluated where high antimicrobial activity is recorded compared to some of the standards.
Experimental
Materials and methods
The sample (skin) for this project was gotten from General Hospital in Abraka, Delta State, Nigeria with sterile containers and was transfer to the laboratory with sterile cellophane. The microbes used for bioactivities of standards (antibiotics), ligands and copper (II) complex were Candida albicans, Aspergillus niger, Klebsiella spp, Shigella spp and Salmonella sp.. Standards (antibiotics) used were purchased from Standard Pharmaceuticals Ltd. All chemicals used were of the analytical reagent grade (AR) and of highest purity available. They included copper (II) sulphate pentahydrate (CuSO4.5H2O), succinic acid and DL-alanine. Methanol and ethanol were used without further purification. De-ionized water was used in all preparations. FT-IR spectrum of the complex was recorded in 400-450 cm-1 range with a Perkin Elmer FT-IR type 1650 spectrophotometer. The complex was run in KBr disc. The UV-V is spectrophotometric measurement was carried out using M501 Single beam UV-Visible spectrophotometer in the range of 200-700 nm at room temperature. The morphology of the complex was examined by scanning electron microscope Joel JEM-5200 at an acceleration voltage of 20 Kv in a magnification range of 30-80 μm. Powder X-ray diffraction (PXRD) studies by PAN analytical X’PERT pro automatic diffractometer operating at 40 kV and 30 mA. The magnetic susceptibility was measured on powdered samples using the Faraday method. The diamagnetic corrections were made by Pascal’s constant and Hg[Co(SCN)4] was used as a calibrant. Molar conductance was measured on an ELICO (CM82T) conductivity bridge. Melting points of the samples were determined on electro thermal capillary apparatus, Gallenkamp, England.
Synthesis of copper(II) Complex
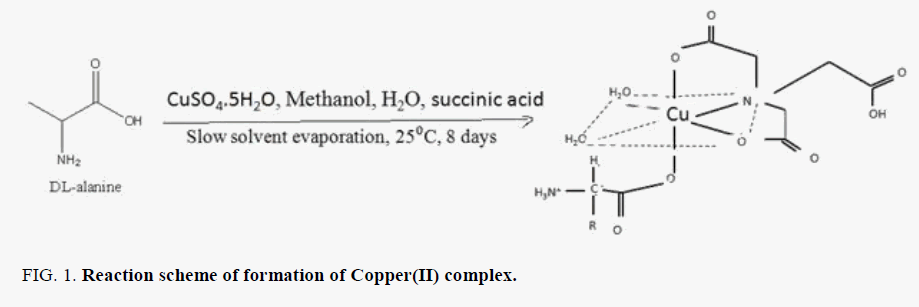
The Copper(II) complex was synthesized with some modification to the method reported in the literature [8]. The copper(II) metal complex was synthesized by self-assembly method under slow solvent evaporation conditions. 1 mmol of succunic acid, 1 mmol of CuSO4.5H2O, and 1 mmol of DL-alanine were dissolved in a mixture of distilled water (10 mL) and methanol (10 mL). The solution was stirred for 2 hours at ambient temperature. The mixture was filtered. The filtrate kept in a cool dry place formed blue crystals on the 8th day. FIG. 1 shows the synthesis of Copper(II) with reactants at 25°C.
Antimicrobial activity tests
The antimicrobial tests were carried out in the Microbiology laboratory of Delta State University, Abraka. Human skin samples was obtained from General Hospital in Abraka, Delta State, Nigeria were used for these tests. The tests were done on five pathogenic micro-organisms, three bacterial strains, Shigella flexneri, Salmonella typhimurium and Klebsella pneumoniae species and two fungi yeasts, Aspergillus niger and Candida albicans. Reference antibacterial and antifungal drugs were evaluated for their antibacterial and antifungal activities on the isolates and their results were compared to those of the synthesized Copper(II) complex and the free ligands.
Preparation of culture media
Sorbitol-MacConkey Agar was prepared according to manufacturer’s directives, autoclaved at 121°C for 15 minutes, allowed to cool to 50°C. The sterilized agar was pour into sterile Petri dishes and the samples were introduced then incubated at 37°C for 24 hours [9].
Standardization of bacteria isolates
The McFarland standard was used for standardization of bacterial isolates. For the preparation of McFarland standard 1% of anhydrous barium chloride (BaCl2) is mixed with 1% (vol/vol) cold pure sulfuric acid (H2SO4). The screw cap was tightly sealed and the bacteria suspension was stored in the dark at room temperature [10].
Results and Discussion
Analytical and physical data
The analytical and physical data, melting point and elemental analysis of synthesized compound and its ligands are tabulated in TABLE 1. The CHNOM calculated values were in a good agreement with the experimental values. The synthesized compound is blue crystalline crystals, stable in air, soluble in DMSO, methanol, and water; partially soluble in ethanol and ammonia. The elemental analyses (TABLE 1) are in agreement with the chemical formula of the Copper(II) compound. Molar conductance value 2.2×103 Ω−1 cm2 mol−1 (1:2:2) shows the synthesized Copper(II) complex is non-electrolytic in nature [11]. Upon complexation process, the pH value of the new complex has been changed compared to their free ligands and free metal ion, which provide an evidence of the successful formation of the metal complex.
| Colour | Physical state | Melting point | % Yield | Conductivity (?-1cm2mol-1) |
pH | %Found(Calc.) | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| C | H | N | O | M | |||||||
| Succinic acid | Colourless | crystalline | 184 | 2.7 | |||||||
| DL alanine | Colourless | powdery | 258 | 2.4 | |||||||
| Copper(II) complex | blue | crystalline | 160-162 | 70.6 | 2.205 | 2.31 | 34.45 (34.67) | 4.88 (4.86) | 8.03 (8.09) | 34.42 (34.23) | 18.8 (18.15) |
Table 1: Analytical and some physical properties of Copper(II) complex and its Ligands
Complex morphology and electronic spectrum of copper(II) complex
The morphology of the complex obtained from scanning electron microscope (SEM) in FIG. 2 shows rectangular sharp edges plate like crystalline structure. The electronic spectrum of Copper(II) complex in methanol recorded in the ultraviolet -visible region shown in FIG. 3 exhibit intra ligand and charge transfer transitions. The spectrum shows a broad band found at 294 nm suggesting octahedral geometry with a unique band due to ²Eg → ²T2g transition [12-14]. The broadness of the band can be taken as an indication of Jahn-Teller distortion from the regular symmetry. The electronic spectrum and the magnetic moment support the stereochemistry of the complex.
Magnetic susceptibility studies
The magnetic moment of the metal complex was calculated from the measured magnetic susceptibility after employing diamagnetic correction. The magnetic moment of the Copper(II) complex was calculated from the expression:
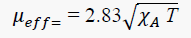
Where χAis the magnetic susceptibility per gram-atom after correction for diamagnetic contributions and T for temperature. The observed magnetic moment of the complex was found to be 1.7 BM which is slightly greater than the spin only value and offers possibility of an octahedral geometry [12].
Infra-red spectra
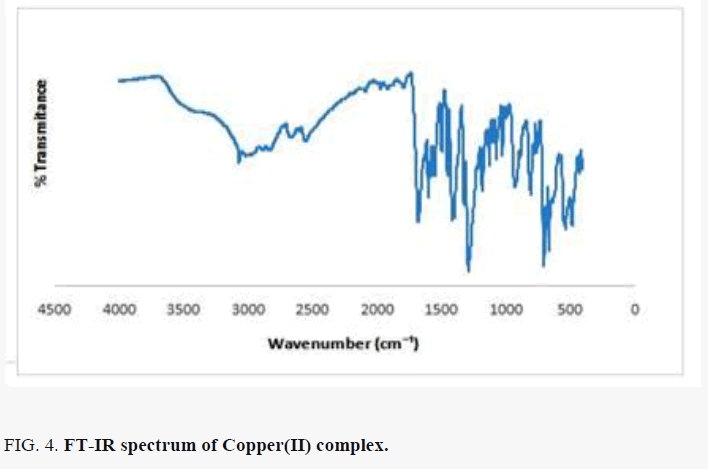
Comparing the FT-IR spectrum provide an insight into the bonding mode of the ligands to the copper(II) metal ion. The FT-IR spectrum of copper(II) complex in FIG. 4 shows some characteristic defined bands and pronounced intensities. The absorption band in the region 3272-3662 cm-l indicates the presence of water in the complex lattice. Band due to C=N in complex is shifted to a lower frequency from 1644 to 1424 cm-1, indicating the coordination of the metal ion to the imino nitrogen. The νasymmetric (COO) band of the free acids at ~1590 cm-1 is shifted to higher wave number in the range 1669- 1596 cm-1 and the νsymmetric (COO) mode observed at ~1400 cm-1 in the spectrum of the amino acids is shifted to lower wave number 1283 cm-1, in the spectrum of the complex, indicating the coordination of the carboxylic acid group via oxygen with the metal ion [14]. The absence of the ν(O-H) band in the copper(II) complex is an indication that, there is loss of proton for O-H group from the free succinic acid and amino acid on coordination revealing those acids are dinegative bidentate ligand coordinating through the carboxylate anion. Some new bands of weak intensity observed in the regions around 662 cm-1 and 485 cm-1 may be ascribed to the M-O and M-N vibrations, respectively [15-17]. It may be noted that these vibration bands are absent in the infra-red spectra of succinic acid as well as DL-alanine.
Powder XRD study
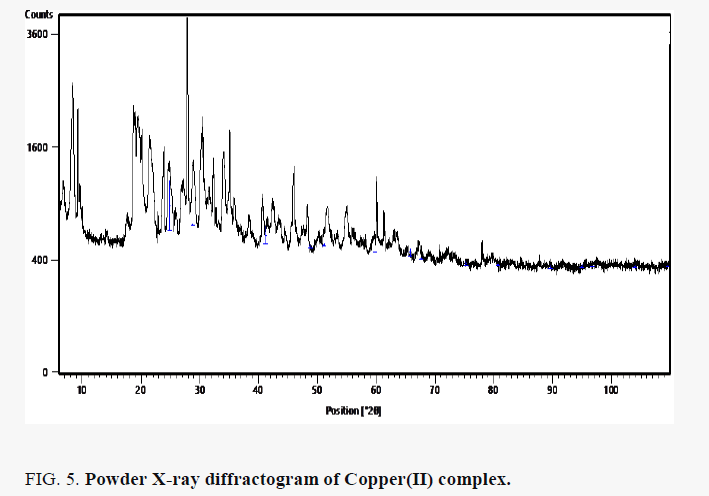
The powder X-ray diffraction pattern of Copper(II) complex presented in FIG. 5 showed that the nanoparticles are crystalline. The reflections in diffraction pattern of Copper(II) complex was recorded with Cu Kα X ray tube in the range 2θ = 4- 100° with λ=1.5406 Å. The unit cell is face-center cubic with lattice parameter of a = 3.542 Å, space group Fm3c. The crystallite size D (2610.773 Å) was estimated from the Scherer’s equation [11]:
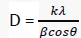
Where D is the crystalline size, K = 0.89 is a correction factor to account for particular changes, β is the full width at half maximum (FWHM) of the most intense diffraction peak, λ = wavelength of X-ray used (Cu K α = 1.5406Å).
Antimicrobial test analysis
The antimicrobial tests were carried out in the Microbiology laboratory of Delta State University, Abraka. The tests were done on five pathogenic micro-organisms, three bacterial strains, Shigella flexneri, Salmonella typhimurium and Klebsiella pneumoniae species and two fungi yeasts, Aspergillus niger and Candida albicans obtained from General Hospital in Abraka, Nigeria. Reference antibacterial and antifungal drugs were evaluated for their antibacterial and antifungal activities on the isolates and their results were compared to those of the free ligands and the Copper(II) complex judged by measuring the diameter of the growth inhibition zone. TABLE 2 represents the isolated organism and biochemical tests employed in identification Biochemical tests and their result were tabulated in TABLE 3 and 4.
| Cultural Characterstics | Catalysis | Citrate | Indole | Motility | Glucose | Lactose | Oxidase | H S2 | Gram rxn | Sucrose | Probable organism |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Cream, | + | - | + | + | + | - | + | - | + | - | Klebsella spp. |
| elongate | |||||||||||
| Cream, | - | - | + | - | - | - | - | - | - | - | Shigella spp |
| elongate | |||||||||||
| Cream, | + | + | - | + | + | - | - | + | - | - | Samonellae |
| ovoid | specie |
Table 2: Biochemical identification and characterization of Bacteria
| 1000 µg | 100 µg | 10 µg | 1 mg | MIC (mg) | |
|---|---|---|---|---|---|
| Klebsella spp. Copper(II) complex | - | - | + | + | 0.1mg |
| Klebsella spp.Succinic acid | - | + | + | + | 1 mg |
| Klebsella spp. DL- Alanine | - | + | + | + | 1 mg |
| Shigellaspp Copper(II) complex | - | + | + | + | 1 mg |
| Shigellaspp Succinic acid | - | + | + | + | 1 mg |
| Shigella spp DL- Alanine | - | + | + | + | 1 mg |
| Samonellae sp. Copper(II) complex | + | + | + | + | >1 mg |
| Samonellae specie Succinic acid | + | + | + | + | >1 mg |
| Samonellae sp, DL- Alanine | - | + | + | + | 1 mg |
Table 3: Minimum inhibitory concentration (MIC) of Copper(II) complex and ligands
| Name of Bacteria Diameter zone of inhibition(mm) | |||
|---|---|---|---|
| Compounds | Diameter zone of inhibition(mm) | ||
| Klebsella Spp | Shigella Spp | Samonellae Specie | |
| Succinic acid | 12.3333 ± 0.33333 | 8.6667 ± 1.33333 | 9.3333 ± 0.66667 |
| DL- Alanine | 10.0000 ± 0.00000 | 9.000 ± 2.08167 | 8.666 ± 0.66667 |
| Copper(II) complex | 18.3333 ± 0.88192 | 17.6667 ± 0.33333 | 20.6667 ± 0.66667 |
| Ciprofloaxacin | 63.3333 ± 48.33333 | 14.6667 ± 0.33333 | 18.0000 ± 1.15470 |
| Norfloxacin | 7.3333 ± 0.66667 | 15.0000 ± 0.57735 | 4.0000 ± 2.00000 |
| Gentamycin | 13.3333 ± 0.66667 | 16.000 ± 0.57735 | 18.6667 ± 0.66667 |
| Amoxil | 6.3333 ± 0.88192 | 12.3333 ± 0.33333 | 14.000 ± 1.15470 |
| Streptomycin | 18.3333 ± 0.88192 | 15.3333 ± 0.33333 | 19.333 ± 0.66667 |
| Ofloxacin | 17.3333 ± 1.33333 | 16.6667 ± 0.66667 | 20.666 ± 0.66667 |
| Erythromycin | 11.3333 ± 0.66667 | 6.000 ± 1.15470 | 11.6667 ± 0.33333 |
| Chloramphenicol | 10.0000 ± 0.00000 | 8.6667 ± 0.66667 | 10.666 ± 0.66667 |
| Ampiclox | 10.6667 ± 0.66667 | 8.6667 ± 0.66667 | 11.3333 ± 0.66667 |
| Levofloaxacin | 12.0000 ± 0.57735 | 12.0000 ± 0.00000 | 14.0000 ± 0.57735 |
Table 4: Results of the antibacterial activity of Copper(II) complex, ligands and standards
Minimum inhibitory concentration (MIC) of the Copper(II) complex and Ligands
TABLE 3 shows the values of Minimum Inhibitory Concentration (MIC) and the minimum bactericidal or fungicidal concentration (MBC or MFC) for each microorganism tested using Copper(II) complex and Ligands. The MIC represents the lowest drug concentration at which a visible inhibition growth of the micro-organisms is observed [17,18]. The MIC values for the different bacteria inoculums incubated at 37°C for 24 hours were determined by using the micro broth dilution test [17]. The lowest MIC was obtained against bacteria Klebsiella spp. at 0.1 mg mL-1 with Copper(II) complex. The MIC values indicate that the Copper(II) complex has a higher activity against the bacteria species than the ligands. Copper(II) complex is more potent and broad spectrum in its activity towards all the tested strains as observed from the MIC values. Comparison of the MIC values of the Copper(II) complex and free ligands with that of the reference drugs indicates that the synthesized complex is potently more active than some of the reference drugs and free ligands.
TABLE 4 is the results of antimicrobial tests carried on Shigella, Salmonella and Klebsiella species The bacterial isolates were sensitive to Copper(II) complex, Ligands and standards. The synthesized compound showed variable enhanced sensitivity pattern of the isolates. The standard drugs, Copper(II) complex, DL- Alanine and succinic acid showed considerable antimicrobial activity against the tested micro-organisms. The Copper(II) complex was observed to have shown favorably enhanced activity against the bacteria and fungi species tested (TABLE 4 and 5). WHONET software package was used for the management of microbiology laboratory data and the analysis of antimicrobial susceptibility test results. The results in TABLE 4 and TABLE 5 show that coordination of the ligands to the metal ion enhanced antimicrobial activity against some of the fungi and bacteria species tested with inhibition zone diameters in the range 16–18 mm. This is in agreement with results in Literature [18-21].
| Zone of inhibition of growth(mm) | Zone of inhibition of growth(mm) | |
|---|---|---|
| Compounds | Aspergillus niger | Candida albicans |
| DL Alanine | 13.0000 ± 0.57735 | 10.6667 ± 0.66667 |
| Succinic acid | 11.3333 ± 0.66667 | 12.6667 ± 0.66667 |
| Copper(II) Complex | 18.00 ± 1.00 | 16.50 ± 0.89 |
| FLU | 16.6667 ± 0.33333 | 14.3333 ± 0.88192 |
| N | 10.6667 ± 0.66667 | 13.3333 ± 0.66667 |
Table 5: Antifungal activity of Copper(II) Complex, Ligands and Standards
Conclusion
Conclusively, The Copper(II) complex and its ligands investigated have potent antifungal effect of being capable of inhibiting the growth of Aspergillus niger and Candida albicans by 50% even at a concentration less than 1 mg (TABLE 3). Copper(II) complex and its ligands show significant antibacterial properties, an indication that they can function as effective antibacterial agents. The copper(II) complex displayed an enhanced potent antifungal effect, suggesting that it may be useful to investigate this agent more fully as a potential antifungal agent in treatment of fungal diseases. This present findings may open a new search for copper(II) complexes of Schiff base ligands to be used in treatment of fungal diseases. Thus, the results presented here suggested that in the future, the copper(II) complexes of Schiff base ligands of DL-alanine and succinic acid have good potential as therapeutic agents.
References
- World Health Organization (WHO) Antimicrobial resistance, 2018.
- Centers for Disease Control and Prevention (CDC) Antibiotic resistance threats in the United States, 2013.
- Smith RA, M?ikanatha NM, Read AF. Antibiotic resistance: a primer and call to action. Health Commun. 2015; 30(3): 309-14.
- Shoukry AA. Complex formation reactions of promethazine copper (II) and various biologically relevant ligands. Synthesis, equilibrium constants, spectroscopic characterization and biological activity. J Solution Chem. 2011; 40(10): 1796-818.
- Hosny NM, El-Dossoki FI. Schiff base complexes derived from 2-acetylpyridine, leucine, and some metal chlorides: their preparation, characterization, and physical properties. J Chem Eng Data. 2008; 53(11): 2567-72.
- Altundas A, Sar? N, Colak N, et al. Synthesis and biological activity of new cycloalkylthiophene-Schiff bases and their Cr (III) and Zn (II) complexes. Med Chem Res. 2010; 19(6): 576-88.
- Lalonde M, Bury W, Karagiaridi O, et al. Transmetalation: Routes to metal exchange within metal?organic frameworks. J Mater Chem. 2013; 1(18): 5453-68.
- Ededet EA, Ayimele GA, Wang ZX, et al. Synthesis, crystal structure and magnetic properties of [Cu (mal)(abpt)(H2O)]. 3/2H2O and [Cu2(sq)(abpt)2]. 2H2O (mal= malonate, sq= squarate, abpt= 4-amino-3, 5-di-2-pyridyl-4H-1, 2, 4 triazole). Bull Chem Soc Ethiop. 2011; 25(1).
- Cheesbrough M. District laboratory practice in tropical countries. Cambridge university press; 2006.
- Sathish K, Chityala VK, Subhashini NJ, et al. Synthesis, Characterization, and Biological and Cytotoxic Studies of Copper (II), Nickel (II), and Zinc (II) Binary Complexes of 3-Amino-5-methyl Isoxazole Schiff Base. ISRN Inorganic Chemistry. 2013.
- Hussain M, Shabeeb GM. Synthesis and Optical Properties of a Biphenyl Compound. Walailak Journal of Science and Technology (WJST). 2013; 11(9): 769-76.
- Reddy SL, Endo T, Reddy GS. Electronic (absorption) spectra of 3d transition metal complexes. In Advanced Aspects of Spectroscopy 2012: 4-48.
- H. Ajayi, N. Hai, B. James. Synthesis and spectra studies and biological activities of Co(II), Cu(II), Ni(II) and Zn(II) complexes with succinic acid. Tur J. Chm.2010, 135: 133-7.
- Kiseleva I, Pyreu D, Krivonogikh T, et al. Thermodynamic study of mixed-ligand complex formation of copper (II) and nickel (II) nitrilotriacetates with amino acids in solution. I. Polyhedron. 2013; 51: 10-7.
- Mahmoud WH, Mohamed GG, El-Dessouky MM. Synthesis, structural characterization, in vitro antimicrobial and anticancer activity studies of ternary metal complexes containing glycine amino acid and the anti-inflammatory drug lornoxicam. J Mol Struct. 2015; 1082: 12-22.
- K. Nakamoto. Infrared and Raman Spectra of Inorganic and Coordination Compounds. 4th edition. New York, NY, USA: John Wiley and Sons; 1986.
- CLSI. Performance Standards for Antimicrobial Disk and Dilution Susceptibility Tests for Bacteria Isolated From Animals. 5th edition. CLSI standard VET01. Wayne, PA: Clinical and Laboratory Standards Institute; 2018.
- EUCAST. The European Committee on Antimicrobial Susceptibility Testing. Breakpoint tables for interpretation of MICs and zone diameters, 2018.
- CLSI. Performance Standards for Antimicrobial Susceptibility Tests. 25th ed. CLSI supplement M100. Wayne, PA: Clinical and Laboratory Standards Institute; 2017.
- Kamaludin NF. Evaluation of antimicrobial activities of organotin (IV) alkylphenyl dithiocarbamate compounds. Asian Journal of Applied Sciences. 2015; 8(2): 165-72.
- Hussain T, Roohi A, Munir S, et al. Biochemical characterization and identification of bacterial strains isolated from drinking water sources of Kohat, Pakistan. Afriscan Journal of Microbiology Research. 2013; 7(16): 1579-90.